4.4.
Training a Unet model to create a synthetic NDVI
The U-Net is a convolutional neural network designed first proposed by (Ronneberger et al., 2015) for image segmentation, recognized by its characteristic symmetric "U"-shaped architecture. It comprises two main paths: a down sampling path that sequentially reduces spatial resolution while extracting features, and an up-sampling path that reconstructs the output to the original image size. The down sampling blocks typically involve 3×3 convolutions and max pooling to halve the image dimensions, while increasing the number of feature channels. The up-sampling path uses transposed convolutions to increase image size, followed by concatenation with feature maps from the corresponding down sampling layer via skip connections. These skip connections are critical for preserving fine-grained spatial details lost during down sampling, leading to more accurate segmentation, particularly at object boundaries.
For the agricultural application being able to delineate the boundaries is particularly important as NDVI inside and outside the field boundaries are primarily different. However, application of U-Net for regression tasks is scarce in the literature. These 2 major motivating factors for the choice of U-Net for NDVI prediction. For prediction of NDVI custom variant of the U-Net architecture was implemented which is also shallower than the original U-Net structure. The shallow U-Net design follows the U-net principles, consisting of an encoder-decoder structure with skip connections, however, reduces the depth of both blocks by reducing the number of convolution blocks to 3 in both encoder and decoder sections.
Encoder
The encoder section consists of a sequence of three convolutional blocks. Each block contains two 2D convolution layers with 3×3 kernels, ReLU activation, and same padding to preserve spatial dimensions. Then a drop out layer with 0.25 rate, followed by a 2×2 max-pooling is applied to reduce spatial resolution. The dropout layer is added to mitigate the overfitting, which is a deviation from original u-net architecture. The number of filters doubles after each convolution block in the encoder section starting from 64 to 256.
Bottleneck
The bottleneck section consists of only one block of 2 convolution layers. As the architecture is 1 step shallower than the original U-net architecture, the filtered in the bottleneck are 512 instead of 1024.
Decoder
Similar to the original U-Net architecture, the decoder section mostly reveres the encoder section using 3 up-sampling blocks to gradually restore spatial resolution. Each up-sampling operation is followed by concatenation with the corresponding encoder feature map (skip connection), allowing high-resolution spatial details to flow directly into the reconstruction path. Each decoder block then applies two convolutional layers (with decreasing filter counts) and a Dropout layer.
For the final output, a Linear activation function is used to predict the continuous NDVI values (a regression output), differing from the original U-Net, which uses a Sigmoid or Softmax activation for segmentation (classification).
The difference between original U-Net structure and the one applied is summed up in the following table-
| Feature | U-Net | Shallow U-Net |
|---|---|---|
| Primary Task | Image Segmentation (Classification) | Image Regression (NDVI Prediction) |
| Depth | 4 Down-sampling/Up-sampling Steps | 3 Down-sampling/Up-sampling Steps |
| Bottleneck Filters | 1024 | 512 |
| Final Activation | Sigmoid | Linear (for continuous regression output) |
The graphical representation of the shallow U-Net is presented below.
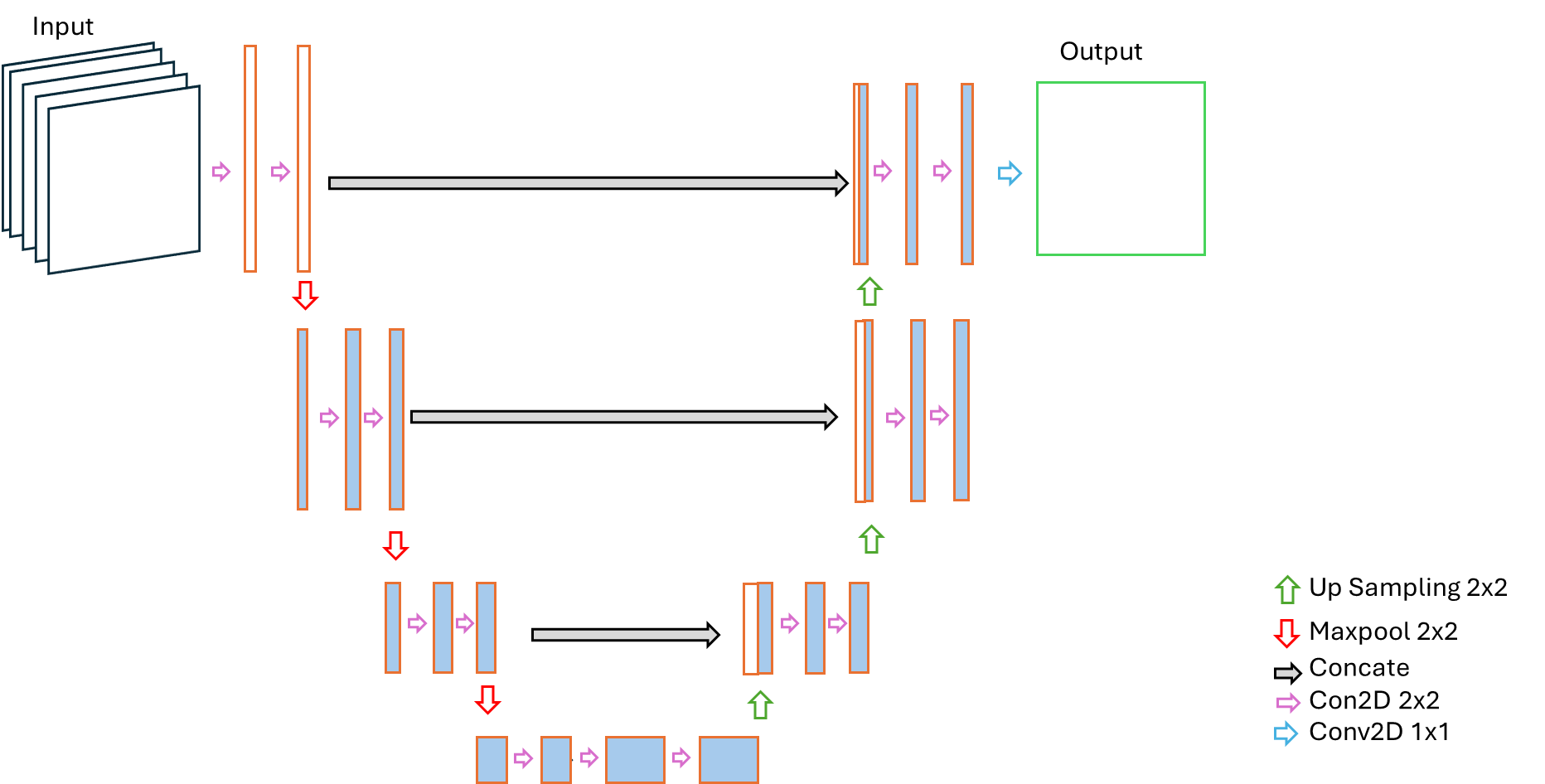
Model evaluation and Comparison with references:
It's essential to monitor model metrics after each epoch to understand a model's performance. Furthermore, comparing these metrics on both the training and validation datasets is crucial for diagnosing whether a model has been overfitted or underfitted. This diagnostic process often involves examining the loss curve, Mean Absolute Error (MAE) curve, and Root Mean Squared Error (RMSE) curve. For this comparison few concepts related to model training should be clarified which are listed below
| Term | Description |
|---|---|
| Model Metrics | Quantifiable measures used to evaluate the performance of a machine learning model, such as accuracy (for classification) or loss (for regression/classification). They help determine how well the model is solving the task. |
| Epoch | One complete pass through the entire training dataset during the model's learning process. An epoch consists of one or more iterations (or batches). |
| Loss Curve | A plot that visualizes the change in the loss function value over each epoch. Monitoring the training loss curve and validation loss curve together helps diagnose overfitting or underfitting. |
| MAE Curve | A plot of the Mean Absolute Error (MAE) metric over each epoch. MAE is the average of the absolute differences between the predicted and actual values. It's less sensitive to outliers than RMSE. |
| RMSE Curve | A plot of the Root Mean Squared Error (RMSE) metric over each epoch. RMSE is the square root of the average of the squared errors. It gives relatively high weight to large errors, making it useful when large errors are particularly undesirable. |
| Overfitting | A phenomenon where a model learns the details and noise in the training data to an extent that it negatively impacts the performance on new, unseen data (like the validation set). This results in excellent performance on the training data but poor performance on the validation data. |
| Underfitting | A phenomenon where a model is too simple to capture the underlying patterns in the data. The model performs poorly on both the training and validation datasets because it hasn't learned the fundamental relationships within the data. |
A plot showing the loss curve, MAE, MSE, and RMSE curves are shown in the below figure, where the red line shows in which epoch of the model training, the best model was found.
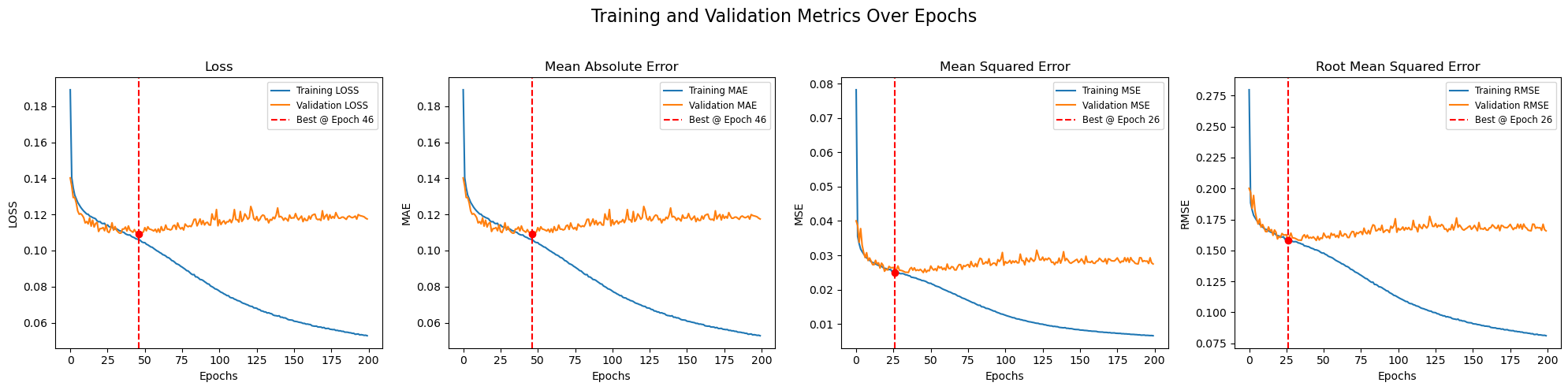
After a best model has been selected with help of metrices, the predictions have to be compared with the visual outputs to validate that the output replicates the real life scenario.
The procedures described in sections 2.3 and 2.4 are applied in the notebook file 1_UNET_TRAIN.ipynb located in the 2_UNET folder.