Section 3
Geospatial data handling; groundwater time series and covariates
Estimated Time: ~30 minutes
Covariates were chosen based on their established impact on the groundwater table head fluctuations (Amanambu et al., 2020; Condon and Maxwell, 2015; Glanville et al., 2023). See section 1 for details
Load required libraries
library(raster)
library(sp)
library(rgdal)
library(dplyr)
library(ggplot2)
library(meteo)
library(rstudioapi)
Define base path and working directory
base_path <- "C:/Users/raza/project"
set.seed(42)
RNGkind(kind = "Mersenne-Twister", normal.kind = "Inversion", sample.kind = "Rejection")
wd <- dirname(rstudioapi::getActiveDocumentContext()$path)
setwd(wd)
Load boundary shapefile and define coordinate systems
utm33 <- "+proj=utm +zone=32 +ellps=WGS84 +datum=WGS84 +units=m +no_defs"
border_wgs84 <- readOGR(paste0(base_path, "/borders/Brandenburg1.shp"))
border <- spTransform(border_wgs84, CRS(utm33))
Create directory for plots if not present
if (!dir.exists("plot")) dir.create("plot")
Define variables
v <- "GWD"
year <- "2001-05"
covariates <- c("GWD", "Lat", "Lon", "SLOPE", "ASP", "DEM",
"LNF", "TPI", "TWI", "TRI", "PREC", "TEMP",
"NDVI", "ET", "EVI")
Convert cleaned CSV data to space-time object (STFDF)
for (year in "2001-05") {
# Load cleaned dataset
file_name <- paste0(base_path, "/clean_", year, ".csv")
bran <- read.csv(file_name)[, -1]
bran <- na.omit(bran)
# Format date
bran$DATE <- as.Date(paste0(bran$DATE, "-01"), format = "%b-%y-%d")
# Extract observation and station information
ob <- bran[, c(5, 6, 10, 18, 19, 20, 21, 22)]
sta <- bran[, c(5, 3, 4, 7, 8, 11, 12, 13, 14, 15, 16, 17)]
sta <- sta[!duplicated(sta), ]
# Create space-time data structure
stfdf <- meteo2STFDF(
obs = ob,
stations = sta,
crs = CRS(utm33),
obs.staid.time = c(1, 2),
stations.staid.lon.lat = c(1, 4, 5)
)
stfdf@sp@data$staid <- 1:nrow(stfdf@sp)
# Save STFDF object for future use
# save(stfdf, file = paste0("stfdf_", year, ".rda"))
}
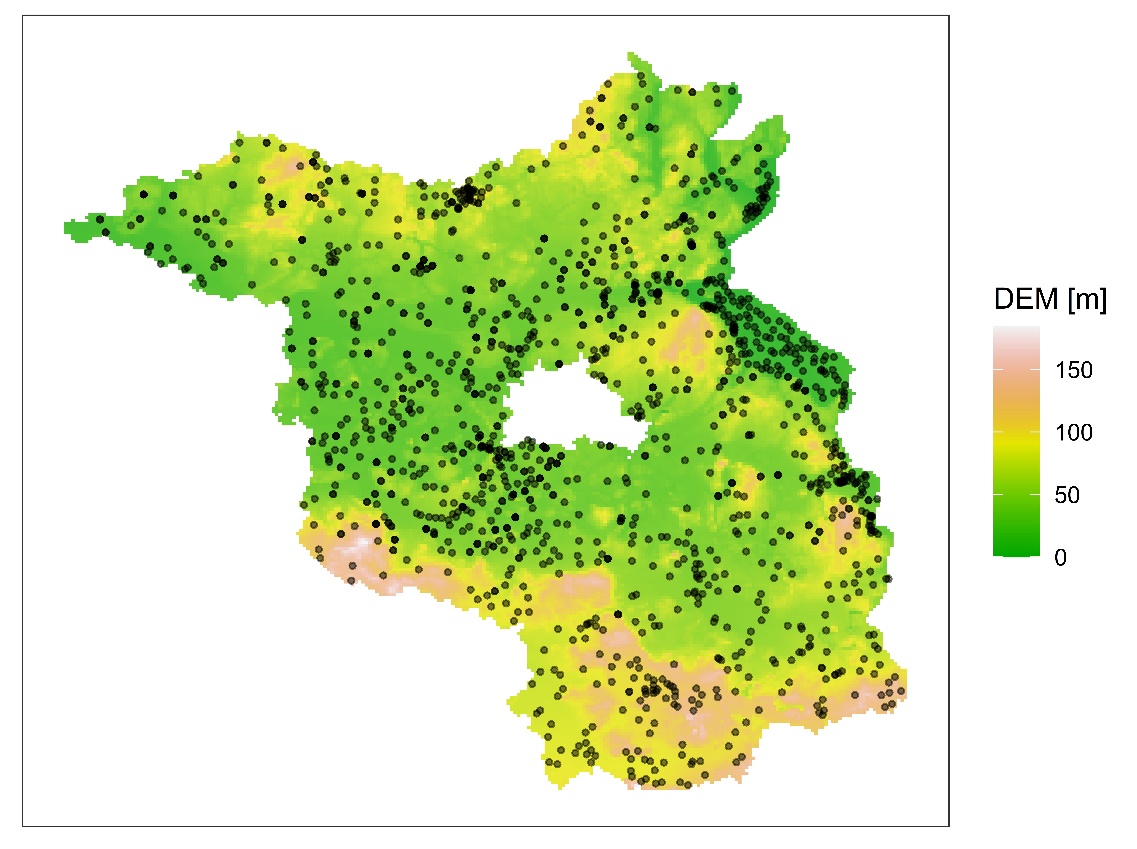
VISUALIZATION 1: STATION LOCATIONS OVER DEM
# Load processed STFDF data
load("stfdf_2001-05.rda")
# Convert spatial projection
stfdf@sp <- spTransform(stfdf@sp, CRS("+proj=utm +zone=31 +ellps=WGS84 +datum=WGS84 +units=m +no_defs"))
# Load and project DEM raster
r <- raster("DEM.tif")
r <- projectRaster(r, crs = CRS("+proj=utm +zone=32 +ellps=WGS84 +datum=WGS84 +units=m +no_defs"))
r <- as(r, "SpatialPixelsDataFrame")
# Create base map with station points
sta_dem_plot <- ggplot() +
geom_raster(data = as.data.frame(r), aes(x = x, y = y, fill = DEM), alpha = 0.8) +
geom_point(data = as.data.frame(stfdf@sp), aes(x = lon, y = lat), size = 0.8, alpha = 0.5) +
scale_fill_gradientn(colours = terrain.colors(100), name = "DEM [m]") +
labs(x = "Longitude", y = "Latitude") +
theme_bw() +
theme(
legend.position = "right",
plot.title = element_text(hjust = 0.5, face = "bold"),
axis.title = element_blank(),
axis.ticks = element_blank(),
axis.text.x = element_blank(),
axis.text.y = element_blank(),
panel.grid = element_blank()
)
# Export figure
jpeg("../plot/stations.jpeg", width = 150, height = 112, units = 'mm', res = 1800)
print(sta_dem_plot)
dev.off()
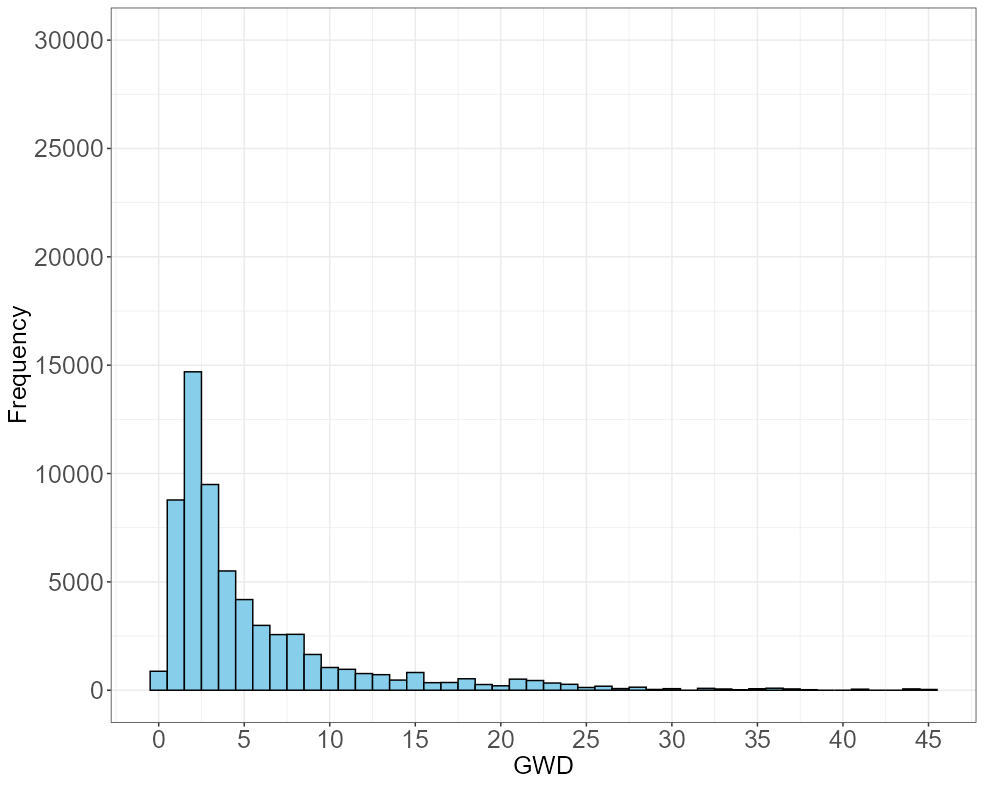
VISUALIZATION 2: HISTOGRAM OF GROUNDWATER DEPTH (GWD)
his <- ggplot(stfdf@data, aes(x = GWD)) +
geom_histogram(binwidth = 1, fill = "skyblue", color = "black") +
labs(x = "GWD", y = "Frequency") +
scale_x_continuous(breaks = seq(0, max(stfdf@data$GWD, na.rm = TRUE), by = 5)) +
scale_y_continuous(limits = c(0, 30000), breaks = seq(0, 30000, by = 5000)) +
theme_bw() +
theme(
plot.title = element_text(size = 18, face = "bold"),
axis.title = element_text(size = 18),
axis.text = element_text(size = 18)
)
# Save histogram
ggsave(filename = "../plot/histogram.jpeg", plot = his, width = 25, height = 20, units = "cm", dpi = 100)