Session 2
The LSP Retrieval Based on S2 and Fused Time Series
In this tutorial we show how to retrieve phenometrics Start-, Peak- and End-of-Season for maize fields in Brandenburg in 2022, using B-spline fitting function and Relative-Amplitude retrieval method. For a pre-defined set of pixel this tutorial allows visual inspection of the fitted B-spline function and extracted DOYs for SoS, PoS and EoS.
Input:
- S2 and Fused NDVI time series
Outputs:
- Output folder named <{current_date}_dataset_smooth_output_v{counter}>
- Retrieval figures visualisation for selected pixels
Steps:
- Global settings
- Definition of loading VI stack for either S2 or Fused NDVI time series
- Definition of plotting and optimising functions
- Processing, retrieval and visualisation for selected pixel set
- Processing full dataset and saving the LSP retrieval results
Main script with basis functions is phenometrics.py.
1. Global settings and Python framework
import os
import numpy as np
import pandas as pd
import concurrent.futures
import rasterio
import json
from scipy.interpolate import splrep, BSpline
from sklearn.linear_model import LinearRegression
from datetime import datetime
from phenometrics import find_max, calculate_relative_amplitude_mean, process_pixel, calculate_sos_eos,plot_ndvi_values, main
import matplotlib.pyplot as plt
import importlib.util# Threshold for starting retrieval doy (needs to be set here for the plotting - the same constant in phenometrics.py)
THRESHOLD_ = 90 2. Loading VI stack for either S2 or Fused NDVI time series
def load_ndvi_stack(directory, filter_threshold=THRESHOLD_):
"""
Load NDVI stack from a directory containing NDVI images.
The threshold is set from the constants if not set manually to an int between 0 and 365.
A two dimensional mask is returned that is true for all pixels that hold at least one value
"""
print('Loading NDVI stack from', directory)
file_paths = sorted([os.path.join(directory, fp) for fp in os.listdir(directory) if fp.endswith('.tif')])
def sort_by_doy(file_paths):
fps = np.array(file_paths)
fp_doys = np.array([os.path.basename(fp).split('_')[-2].split('.')[0] for fp in fps])
sorted_indices = np.argsort(fp_doys)
return fps[sorted_indices]
file_paths = sort_by_doy(file_paths)
doys = np.array([int(os.path.basename(fp).split('_')[-2].split('.')[0]) for fp in file_paths], dtype=np.int64)
if filter_threshold is not None:
file_paths = [fp for fp in file_paths if int(os.path.basename(fp).split('_')[-2].split('.')[0]) >= filter_threshold]
doys = np.array([int(os.path.basename(fp).split('_')[-2].split('.')[0]) for fp in file_paths], dtype=np.int64)
ndvi_stack = []
mask_stack = []
for fp in file_paths:
with rasterio.open(fp) as dataset:
ndvi_data = dataset.read(1).astype(np.float64)
ndvi_stack.append(ndvi_data[LOAD_MIN_ROW:LOAD_MAX_ROW, LOAD_MIN_COL:LOAD_MAX_COL])
# ndvi_stack.append(ndvi_data)
mask_stack.append(np.isnan(ndvi_data[LOAD_MIN_ROW:LOAD_MAX_ROW, LOAD_MIN_COL:LOAD_MAX_COL]))
with rasterio.open(file_paths[0]) as dataset:
meta = dataset.meta
ndvi_stack = np.array(ndvi_stack)
mask_stack = np.array(mask_stack)
valid_image_area_mask = ~np.any(mask_stack, axis=0)
if len(doys) < 3:
raise ValueError(f'Not enough images in the stack (less than 3!) in directory {directory} with threshold {filter_threshold}')
return ndvi_stack, doys, valid_image_area_mask, meta3. Plotting and optimising functions
def convert_to_serializable(obj):
if obj is np.nan:
return None
if isinstance(obj, np.generic):
return obj.item()
return objdef serialize_sos_eos_dict(sos_eos_dict):
serialized_dict = {
'pixel_idx': sos_eos_dict['pixel_idx'],
'row': sos_eos_dict['row'],
'col': sos_eos_dict['col'],
'smooth': sos_eos_dict['smooth'],
'threshold_start': sos_eos_dict['threshold_start'],
'threshold_end': sos_eos_dict['threshold_end'],
'org_ndvi_doys': list(convert_to_serializable(sos_eos_dict['org_ndvi_doys'])),
'org_ndvi_values': list(convert_to_serializable(sos_eos_dict['org_ndvi_values'])),
'sos_median_of_slope_doy': convert_to_serializable(sos_eos_dict['sos_median_of_slope']),
'eos_median_of_slope_doy': convert_to_serializable(sos_eos_dict['eos_median_of_slope']),
'sos_doys': list(convert_to_serializable(sos_eos_dict['sos_doys'])) if np.any(sos_eos_dict['sos_doys']) else None,
'eos_doys': list(convert_to_serializable(sos_eos_dict['eos_doys'])) if np.any(sos_eos_dict['eos_doys']) else None,
'sos_ndvi_values': list(convert_to_serializable(sos_eos_dict['sos_ndvi_values'])) if np.any(sos_eos_dict['sos_ndvi_values']) else None,
'eos_ndvi_values': list(convert_to_serializable(sos_eos_dict['eos_ndvi_values'])) if np.any(sos_eos_dict['eos_ndvi_values']) else None,
'amplitude': convert_to_serializable(sos_eos_dict['amplitude']) if np.any(sos_eos_dict['amplitude']) else None,
'max_ndvi_doy': convert_to_serializable(sos_eos_dict['max_ndvi_doy']),
'fine_doys': list(convert_to_serializable(sos_eos_dict['fine_doys'])),
'base': convert_to_serializable(sos_eos_dict['base']),
'relative_amplitude_mean': convert_to_serializable(sos_eos_dict['relative_amplitude_mean_ndvi']),
'sos_relative_amplitude_mean': convert_to_serializable(sos_eos_dict['sos_relative_amplitude_mean_ndvi']),
'eos_relative_amplitude_mean': convert_to_serializable(sos_eos_dict['eos_relative_amplitude_mean_ndvi']),
'sos_relative_amplitude_mean_doy': convert_to_serializable(sos_eos_dict['sos_relative_amplitude_mean']),
'eos_relative_amplitude_mean_doy': convert_to_serializable(sos_eos_dict['eos_relative_amplitude_mean']),
}
return serialized_dict4. Processing, retrieval and visualisation for selected pixel set
def process_stack(input_dir):
"""Main function to process NDVI data."""
print(f'Start processing {input_dir}')
start = datetime.now()
current_date = datetime.now().date().isoformat()
ndvi_stack, doys, mask, meta = load_ndvi_stack(input_dir)
num_rows, num_cols = ndvi_stack[0].shape
for row in range(num_rows):
for col in range(num_cols):
if mask[row, col]:
# try:
sos_eos_dict = process_pixel(row, col, ndvi_stack[:, row, col], doys, num_cols)
sos_eos_dict['row'] = LOAD_MIN_ROW + row
sos_eos_dict['col'] = LOAD_MIN_COL + col
sos_eos_dict.update({
'org_ndvi_doys': doys,
'org_ndvi_values': ndvi_stack[:, row, col],
})
# print('sos_eos_dict', sos_eos_dict)
serialized_dict = serialize_sos_eos_dict(sos_eos_dict)
plot_ndvi_values(serialized_dict)
# Pixels set for visualisation available for 2021
LOAD_MIN_COL = 143
LOAD_MAX_COL = LOAD_MIN_COL + 1
LOAD_MIN_ROW = 190
LOAD_MAX_ROW = 191process_stack('./Dataset/S2/2021/')
process_stack('./Dataset/Fused/2021/')Start processing ./Dataset/S2/2021/
Loading NDVI stack from ./Dataset/S2/2021/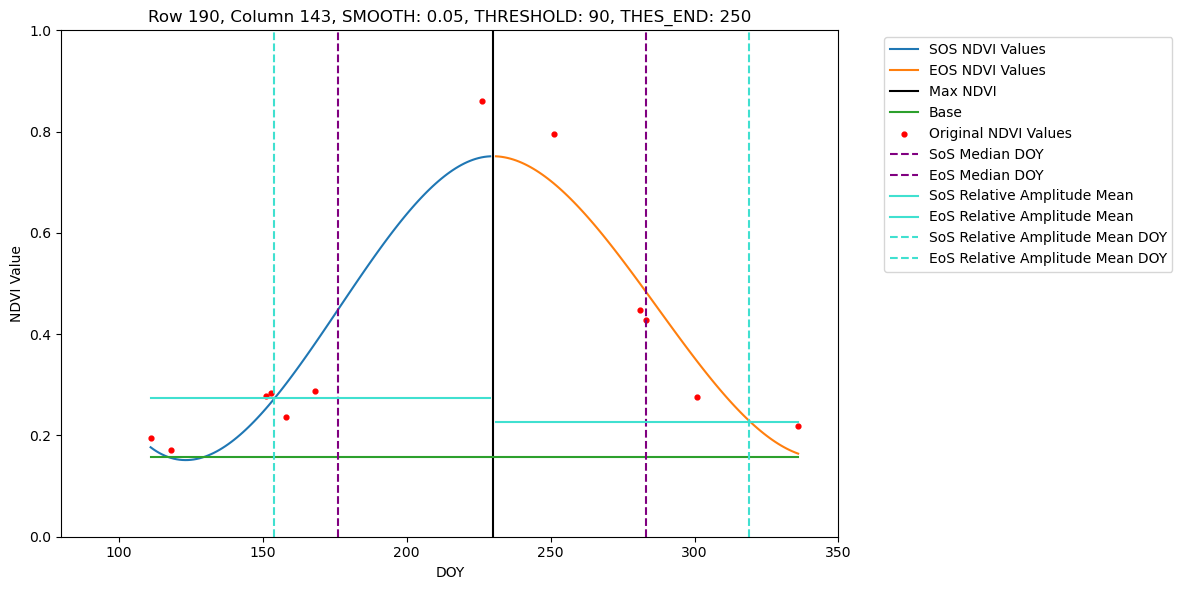
Start processing ./Dataset/Fused/2021/
Loading NDVI stack from ./Dataset/Fused/2021/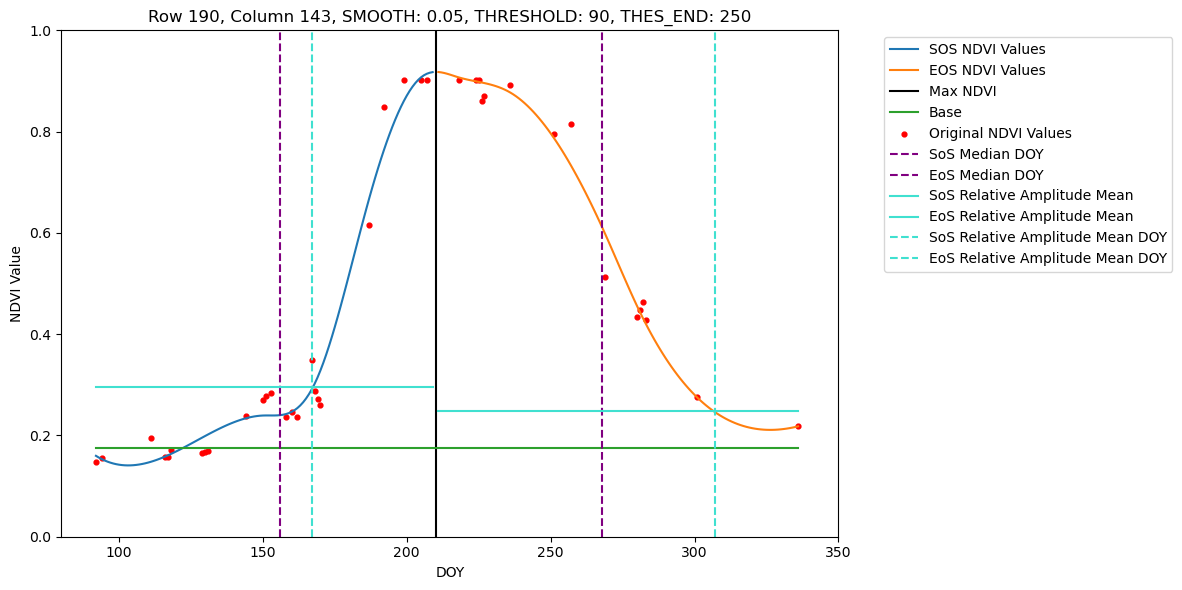
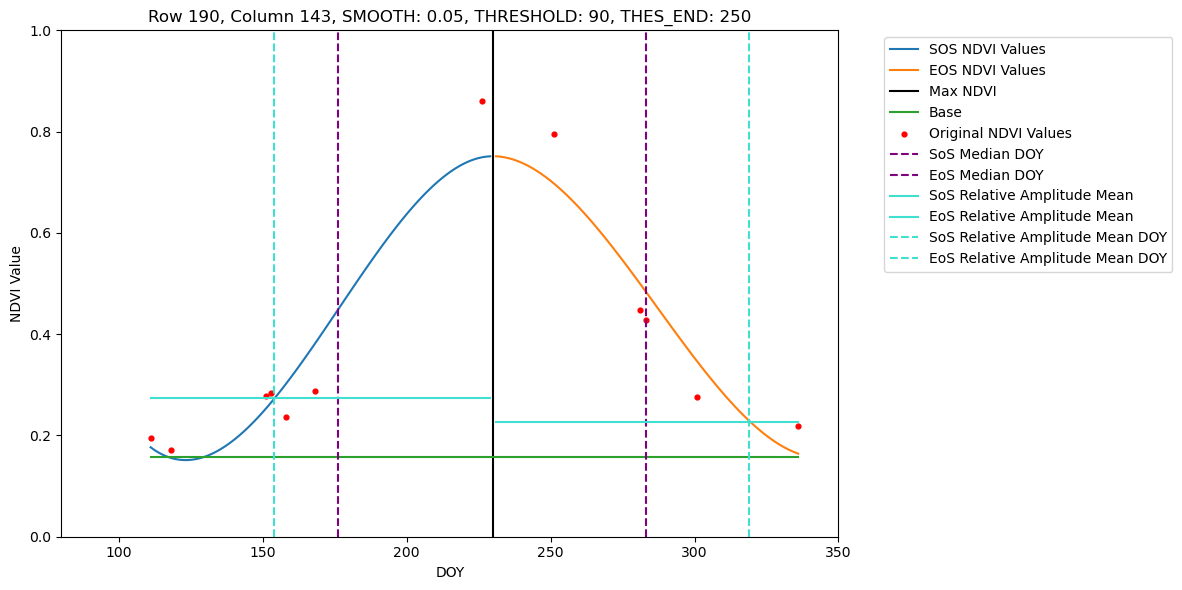
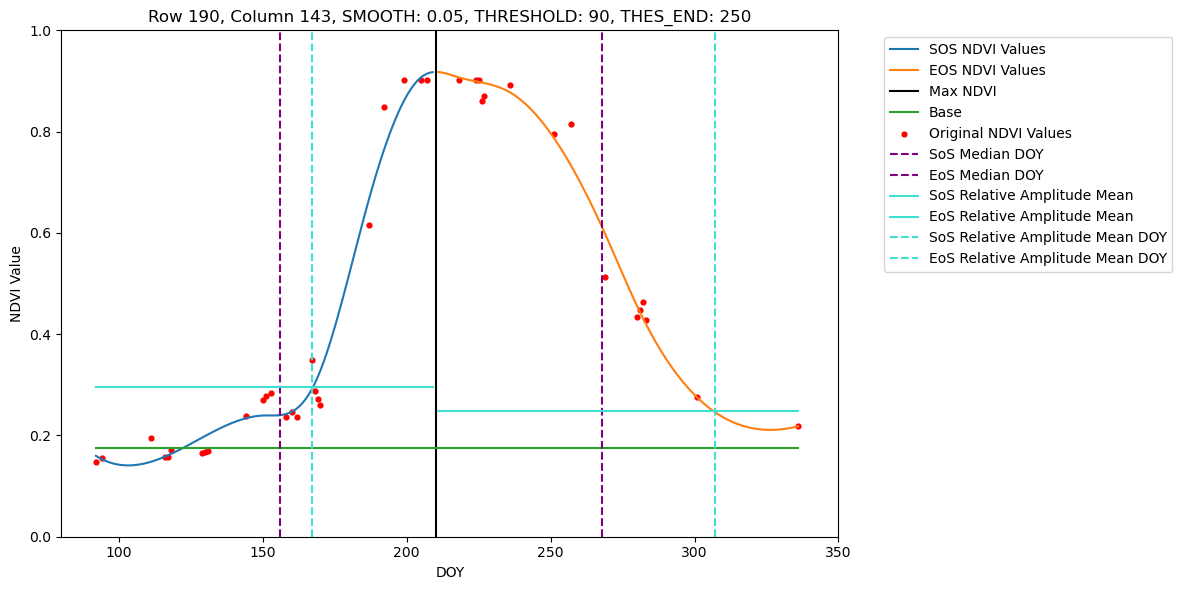
# Pixels set for visualisation available for 2022
LOAD_MIN_COL = 234
LOAD_MAX_COL = LOAD_MIN_COL + 1
LOAD_MIN_ROW = 302
LOAD_MAX_ROW = 303process_stack('./Dataset/S2/2022/')
process_stack('./Dataset/Fused/2022/')Start processing ./Dataset/S2/2022/
Loading NDVI stack from ./Dataset/S2/2022/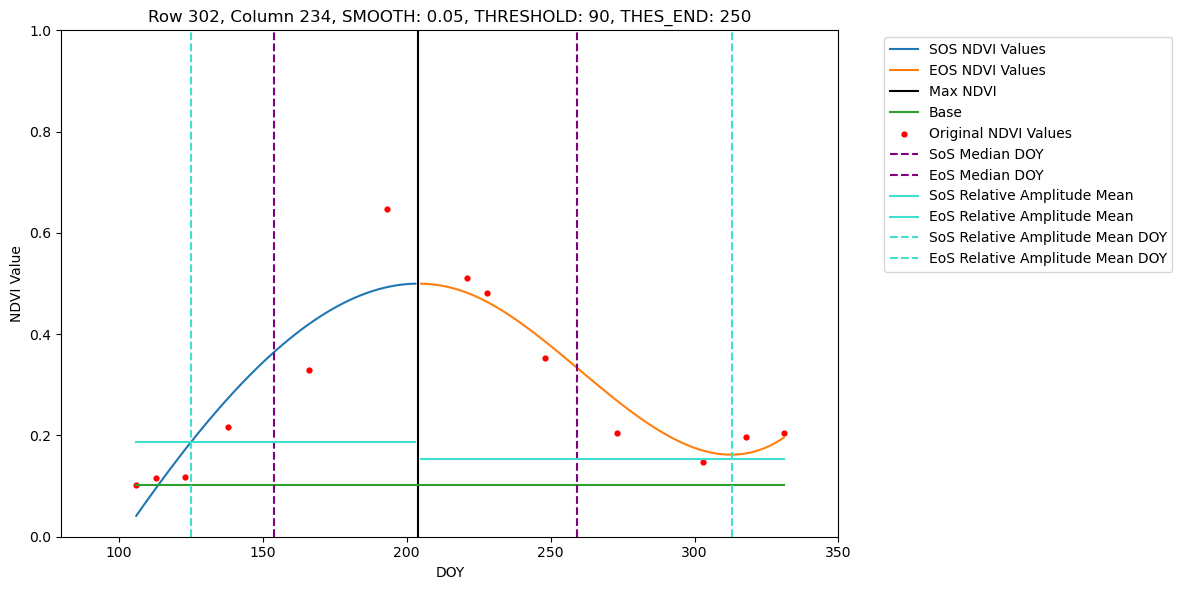
Start processing ./Dataset/Fused/2022/
Loading NDVI stack from ./Dataset/Fused/2022/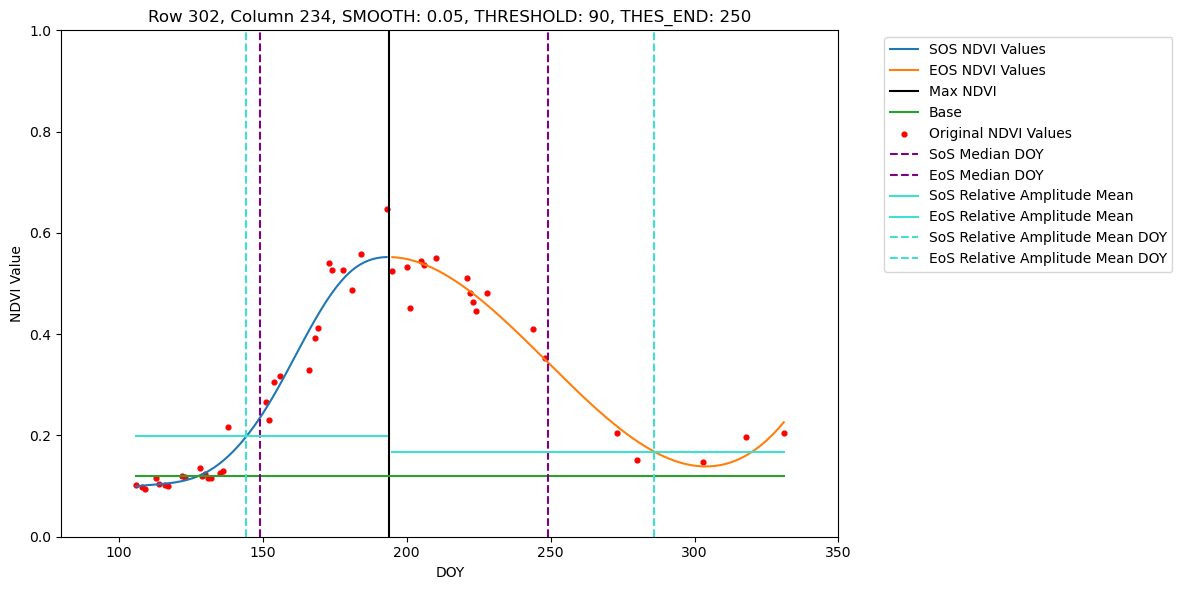
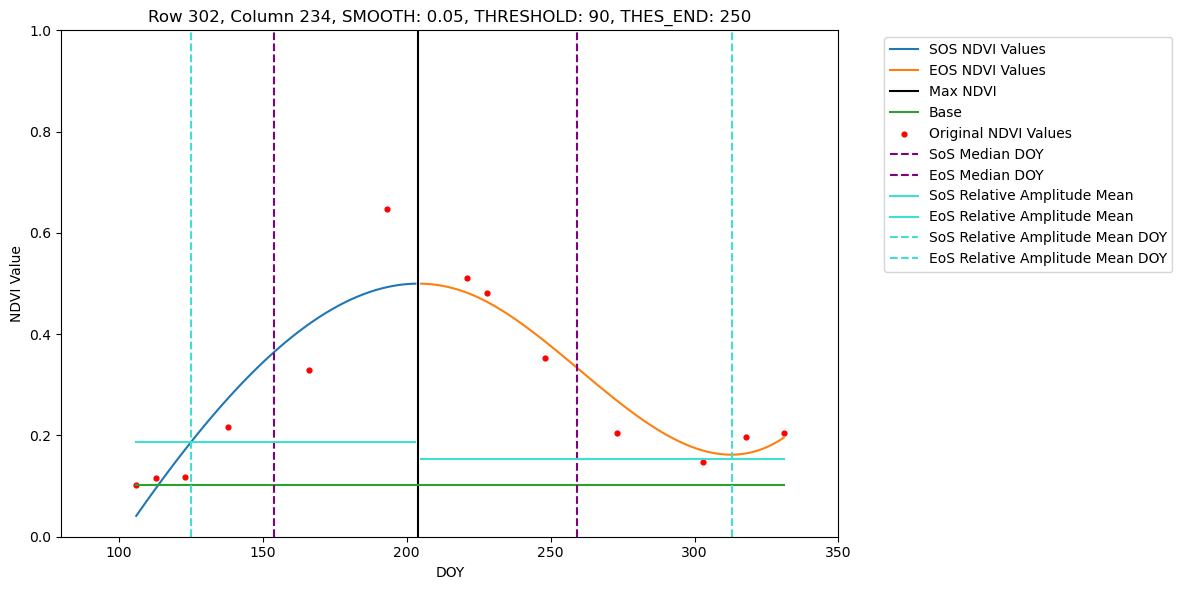
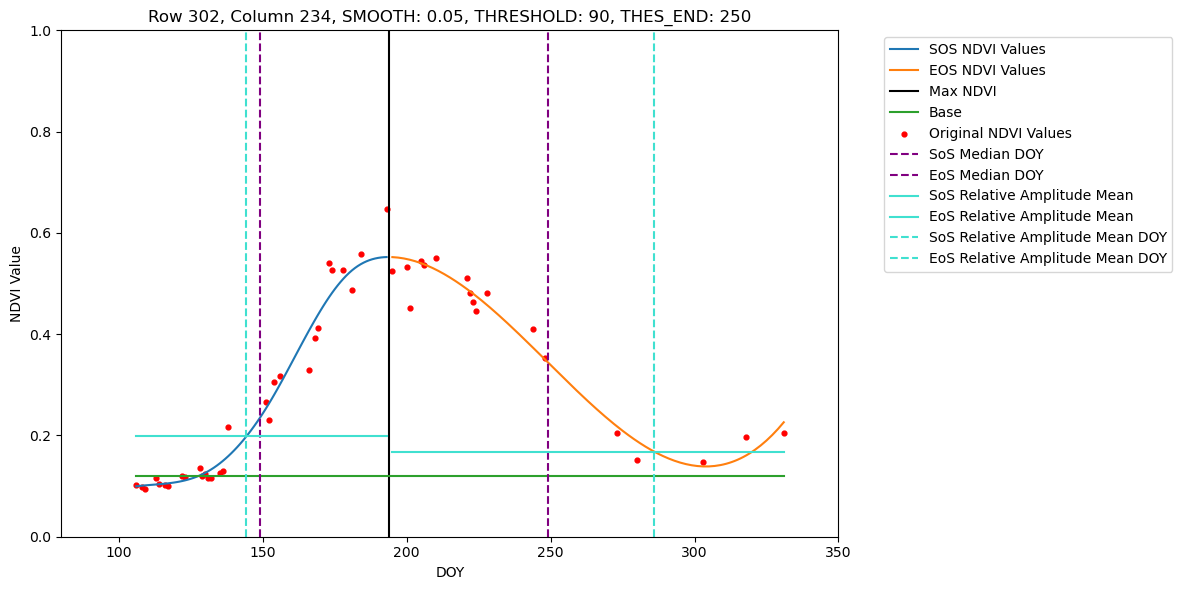
5. Processing full dataset and saving retrieval results
# Processing Sentinel-2 and Fused time series 2021
main('./Dataset/S2/2021/')
main('./Dataset/Fused/2021/')Start processing ./Dataset/S2/2021/
Loading NDVI stack from ./Dataset/S2/2021/
./Dataset/S2/2021/ (397, 558)
writing ./Dataset/S2\2025-09-09_S2_2021_0.05_output_v1\S2_max_2021_ndvi_doy.tif
writing ./Dataset/S2\2025-09-09_S2_2021_0.05_output_v1\S2_sos_2021_median_of_slope.tif
writing ./Dataset/S2\2025-09-09_S2_2021_0.05_output_v1\S2_sos_2021_relative_amplitude_mean.tif
writing ./Dataset/S2\2025-09-09_S2_2021_0.05_output_v1\S2_eos_2021_median_of_slope.tif
writing ./Dataset/S2\2025-09-09_S2_2021_0.05_output_v1\S2_eos_2021_relative_amplitude_mean.tif
Processing ./Dataset/S2/2021/ took 0:00:20.807563
Processing of the directory ./Dataset/S2/2021/ was completed in 0:00:20.807563.
Start processing ./Dataset/Fused/2021/
Loading NDVI stack from ./Dataset/Fused/2021/
./Dataset/Fused/2021/ (397, 558)
writing ./Dataset/Fused\2025-09-09_Fused_2021_0.05_output_v1\Fused_max_2021_ndvi_doy.tif
writing ./Dataset/Fused\2025-09-09_Fused_2021_0.05_output_v1\Fused_sos_2021_median_of_slope.tif
writing ./Dataset/Fused\2025-09-09_Fused_2021_0.05_output_v1\Fused_sos_2021_relative_amplitude_mean.tif
writing ./Dataset/Fused\2025-09-09_Fused_2021_0.05_output_v1\Fused_eos_2021_median_of_slope.tif
writing ./Dataset/Fused\2025-09-09_Fused_2021_0.05_output_v1\Fused_eos_2021_relative_amplitude_mean.tif
Processing ./Dataset/Fused/2021/ took 0:00:28.194966
Processing of the directory ./Dataset/Fused/2021/ was completed in 0:00:28.194966.# Processing Sentinel-2 and Fused time series 2022
main('./Dataset/S2/2022/')
main('./Dataset/Fused/2022/')Start processing ./Dataset/S2/2022/
Loading NDVI stack from ./Dataset/S2/2022/
./Dataset/S2/2022/ (371, 452)
writing ./Dataset/S2\2025-09-09_S2_2022_0.05_output_v1\S2_max_2022_ndvi_doy.tif
writing ./Dataset/S2\2025-09-09_S2_2022_0.05_output_v1\S2_sos_2022_median_of_slope.tif
writing ./Dataset/S2\2025-09-09_S2_2022_0.05_output_v1\S2_sos_2022_relative_amplitude_mean.tif
writing ./Dataset/S2\2025-09-09_S2_2022_0.05_output_v1\S2_eos_2022_median_of_slope.tif
writing ./Dataset/S2\2025-09-09_S2_2022_0.05_output_v1\S2_eos_2022_relative_amplitude_mean.tif
Processing ./Dataset/S2/2022/ took 0:00:17.690906
Processing of the directory ./Dataset/S2/2022/ was completed in 0:00:17.690906.
Start processing ./Dataset/Fused/2022/
Loading NDVI stack from ./Dataset/Fused/2022/
./Dataset/Fused/2022/ (371, 452)
writing ./Dataset/Fused\2025-09-09_Fused_2022_0.05_output_v1\Fused_max_2022_ndvi_doy.tif
writing ./Dataset/Fused\2025-09-09_Fused_2022_0.05_output_v1\Fused_sos_2022_median_of_slope.tif
writing ./Dataset/Fused\2025-09-09_Fused_2022_0.05_output_v1\Fused_sos_2022_relative_amplitude_mean.tif
writing ./Dataset/Fused\2025-09-09_Fused_2022_0.05_output_v1\Fused_eos_2022_median_of_slope.tif
writing ./Dataset/Fused\2025-09-09_Fused_2022_0.05_output_v1\Fused_eos_2022_relative_amplitude_mean.tif
Processing ./Dataset/Fused/2022/ took 0:00:22.045057
Processing of the directory ./Dataset/Fused/2022/ was completed in 0:00:22.045057.