Section 6
Conventional ML – RFSI
Estimated Time: ~25 minutes
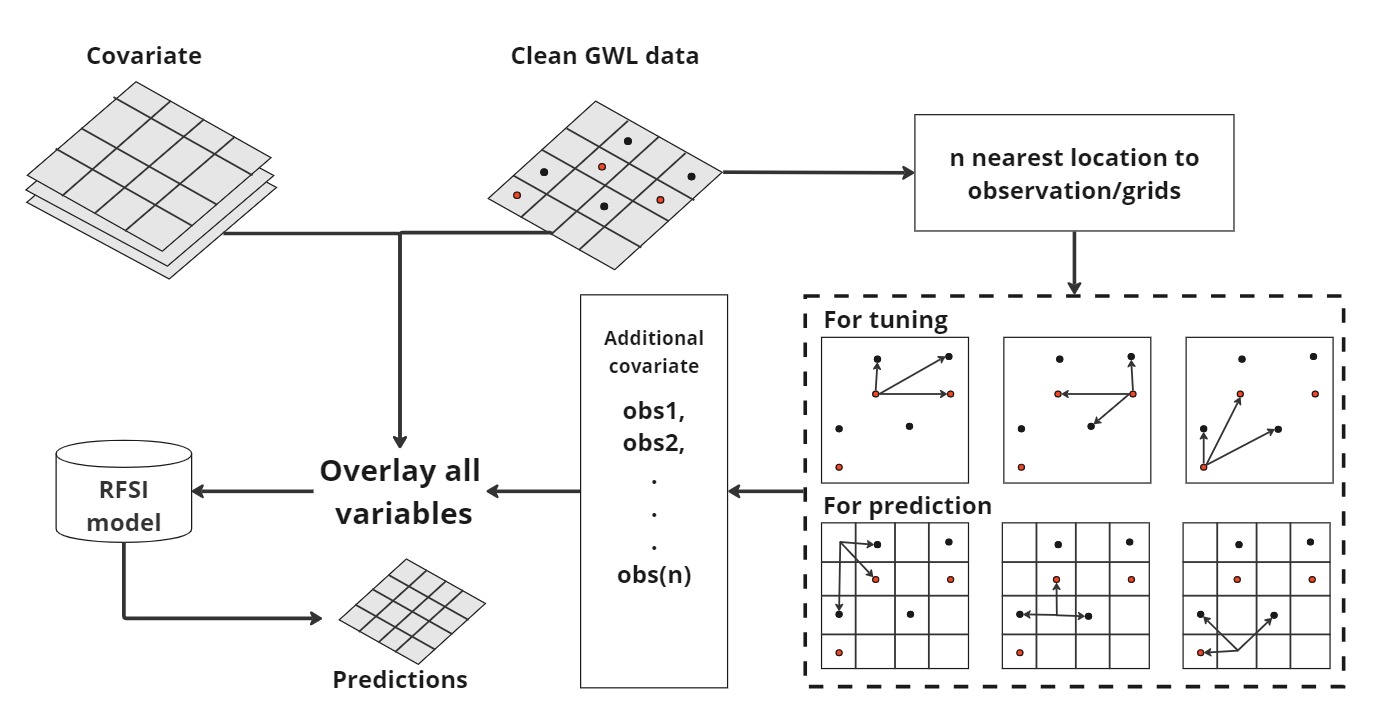
In this section, we apply the Random Forest Spatial Interpolation (RFSI) method to improve groundwater level predictions. Standard Random Forest models do not explicitly account for spatial autocorrelation, except indirectly through spatially correlated covariates. Since nearby observations provide valuable information for estimating values at unsampled locations, we enhance the model by including additional spatial covariates. These covariates represent the observed values at the n nearest locations and the corresponding distances between these locations and each prediction point (Fig. 1)
Inputs:
- Grid based MODIS ET, NDVI, EVI values
- Temperature and precipitation data from German weather station (DWD)
- Coordinates
- observations at n nearest locations
- distance of each observation from these locations
Output
- Trained RFsp model
- Write trained model predictions on the testing data set.
- Tuned set of model parameters
- Computational time
Section 6
Conventional ML – RFSI
####################################
setwd("C:/Users/raza/project")
base_path <- "C:/Users/raza/project/plot"
year_ranges <- c("2001-05")
# Loop through years from 2001 to 2005
for (year in year_ranges) {
start_time <- Sys.time()
cat("Processing year:", year, "\n")
file_name <- paste0("stfdf_", year, ".rda")
# Load spatiotemporal data frame for the current year
load(file = file_name)
# Convert the data to a format suitable for modeling
GWD_df <- as.data.frame(stfdf)
covariates <- c("GWD", "Lat", "Lon", "SLOPE", "ASP", "DEM", "LNF", "TPI", "TWI", "TRI", "PREC", "TEMP", "NDVI", "ET", "EVI")
GWD_df <- GWD_df[, c("Lon", "Lat", "sp.ID", "time", covariates)]
GWD_df = GWD_df[complete.cases(GWD_df), ]
time = sort(unique(GWD_df$time))
daysNum = length(time)
days = gsub("-", "", time, fixed=TRUE)
nos <- 6:14
min.node.size <- 5:6
#min.node.size <- 4:6 # 2:6
sample.fraction <- seq(1, 0.632, -0.05)
ntree <- 250
n_obs = max(nos)
#mtry <- 3:(9 + 2 * n_obs)
mtry <- 3
indices <- CreateSpacetimeFolds(GWD_df, spacevar = "sp.ID", k = 5, seed = 42)
hp <- expand.grid(min.node.size = min.node.size, mtry = mtry, no = nos, sf = sample.fraction)
hp <- hp[hp$mtry < (9 + 2 * hp$no - 1), ]
hp <- hp[sample(nrow(hp), 100), ]
hp <- hp[order(hp$no), ]
rmse_hp <- rep(NA, nrow(hp))
lowest_rmse <- Inf # Initialize with a high value
iteration_since_lowest <- 0 # Counter for iterations since the lowest RMSE
for (h in 1:nrow(hp)) {
comb <- hp[h, ]
print(paste("combination: ", h, sep = ""))
print(comb)
dev_df <- GWD_df
fold_obs <- c()
fold_pred <- c()
for (f in 1:length(indices$index)) {
print(paste("fold: ", f, sep = ""))
cols <- c(covariates, paste("dist", 1:(comb$no), sep = ""), paste("obs", 1:(comb$no), sep = ""))
dev_df1 <- dev_df[indices$index[[f]], ]
cpus <- detectCores() - 1
registerDoParallel(cores = cpus)
nearest_obs <- foreach (t = time) %dopar% {
dev_day_df <- dev_df1[as.numeric(dev_df1$time) == t, c("Lon", "Lat", "GWD")]
day_df <- dev_df[as.numeric(dev_df$time) == as.numeric(t), c("Lon", "Lat", "GWD")]
if (nrow(day_df) == 0) {
return(NULL)
}
return(meteo::near.obs(
locations = day_df,
observations = dev_day_df,
obs.col = "GWD",
n.obs = comb$no
))
}
stopImplicitCluster()
nearest_obs <- do.call("rbind", nearest_obs)
dev_df <- cbind(dev_df, nearest_obs)
dev_df = dev_df[complete.cases(dev_df), ]
dev_df1 <- dev_df[indices$index[[f]], cols]
dev_df1 <- dev_df1[complete.cases(dev_df1), ]
val_df1 <- dev_df[indices$indexOut[[f]], cols]
val_df1 <- val_df1[complete.cases(val_df1), ]
model <- ranger(GWD ~ ., data = dev_df1, importance = "none", seed = 42,
num.trees = ntree, mtry = comb$mtry,
splitrule = "variance", # variance extratrees
min.node.size = comb$min.node.size,
sample.fraction = comb$sf,
oob.error = FALSE)
fold_obs <- c(fold_obs, val_df1$GWD)
fold_pred <- c(fold_pred, predict(model, val_df1)$predictions)
}
rmse_hp[h] <- sqrt(mean((fold_obs - fold_pred)^2, na.rm = TRUE))
print(paste("rmse: ", rmse_hp[h], sep = ""))
if (rmse_hp[h] < lowest_rmse) {
lowest_rmse <- rmse_hp[h]
iteration_since_lowest <- 0 # Reset counter
print(paste("New lowest RMSE:", lowest_rmse))
} else {
iteration_since_lowest <- iteration_since_lowest + 1
print(paste("Iterations since lowest RMSE:", iteration_since_lowest))
}
# Stop if 5 iterations have passed since the lowest RMSE was found
if (iteration_since_lowest >= 5) {
print(paste("Stopping after", iteration_since_lowest, "iterations since the lowest RMSE"))
break
}
}
dev_parameters <- hp[which.min(rmse_hp), ]
print(dev_parameters)
cpus <- detectCores() - 1
registerDoParallel(cores = cpus)
nearest_obs <- foreach (t = time) %dopar% {
day_df <- GWD_df[GWD_df$time == t, c("Lon", "Lat", "GWD")]
if (nrow(day_df) == 0) {
return(NULL)
}
return(near.obs(
locations = day_df,
observations = day_df,
obs.col = "GWD",
n.obs = dev_parameters$no
))
}
stopImplicitCluster()
nearest_obs <- do.call("rbind", nearest_obs)
GWD_df <- cbind(GWD_df, nearest_obs)
cols <- c(covariates, paste("dist", 1:(dev_parameters$no), sep = ""), paste("obs", 1:(dev_parameters$no), sep = ""))
GWD_df <- GWD_df[, cols]
colnames(GWD_df)
a <- GWD_df
# Identify unique combinations of latitude and longitude
unique_coords <- unique(GWD_df[, c("Lat", "Lon")])
# Set a seed for reproducibility
set.seed(123)
# Create an index for splitting the data
index <- createDataPartition(1:nrow(unique_coords), p = 0.8, list = FALSE)
# Use the index to split the unique coordinates into training and testing sets
train_coords <- unique_coords[index, ]
test_coords <- unique_coords[-index, ]
# Merge the original data with the training and testing coordinates
train_data <- merge(GWD_df, train_coords, by = c("Lat", "Lon"))
test_data <- merge(GWD_df, test_coords, by = c("Lat", "Lon"))
nrow(test_data)
write_csv(test_data, "test_data_lat_lon.csv")
# Select covariates
covariates <- colnames(GWD_df)
covariates <- covariates[covariates != "GWD"] # Assuming "GWD" is the response variable
# Subset the data frames with selected covariates
train_data <- train_data[, c("GWD", covariates)]
test_data <- test_data[, c("GWD", covariates)]
rfsi_model <- ranger(GWD ~ ., data = train_data, importance = "impurity", seed = 42,
num.trees = ntree, mtry = dev_parameters$mtry,
splitrule = "extratrees",
min.node.size = dev_parameters$min.node.size,
sample.fraction = dev_parameters$sf,
quantreg = TRUE) ### quantreg???
# save(rfsi_model, file = "../models/RFSI.rda")
save(rfsi_model, file = paste0("../models/RFSI_", year, ".rda"))
# Create and save plot_data
plot_data <- data.frame(Lat = train_data$Lat, Lon = train_data$Lon, Observations = train_data$GWD, Predictions = rfsi_model$predictions)
plot_file <- paste0(base_path, (paste0("\\model_training\\rfsi_training_", year, ".csv")))
write.csv(plot_data, plot_file)
# Make predictions on the test data using the trained model
predictions <- predict(rfsi_model, data = test_data, type = "response")
# Add the predictions to the test_data dataframe
test_data$predicted_GWD <- predictions$predictions
#test_data <- test_data[, c("GWD", "predicted_GWD")]
plot_data_test <- data.frame(Lat = test_data$Lat, Lon = test_data$Lon, GWD = test_data$GWD, predicted_GWD = test_data$predicted_GWD)
# Save test_data with predictions as a CSV file
test_file <- paste0(base_path, (paste0("\\model_testing\\rfsi_test_", year, ".csv")))
write.csv(plot_data_test, test_file)
end_time <- Sys.time()
computation_time <- end_time - start_time
#cat("dev_parameters for year", year, ":", dev_parameters, "\n")
cat("Computation time for year", year, ":", computation_time, "\n")
# Save dev_parameters, RMSE, and computation_time to a text file
result_file <- paste0(base_path, "\\computation_times_rfsi.txt")
write(paste("Year:", year), file = result_file, append = TRUE)
write(paste("dev_parameters:", toString(dev_parameters)), file = result_file, append = TRUE)
write(paste("RMSE:", min(rmse_hp, na.rm = TRUE)), file = result_file, append = TRUE)
write(paste("Computation time:", computation_time), file = result_file, append = TRUE)
write("\n", file = result_file, append = TRUE)
}