Session 1
Training a U-Net Network
In this tutorial we will see how to train a U-Net model from scratch for predicting NDVI from Sentinel 1 Images and Validate the predictions with the ground truth NDVIS generated from Sentil 2 Optical data.
Input:
- Train, Test, and Validation CSVs
- Sentinel 1 Images and Meta Data Outputs:
- Trained Unet Model
- Precited NDVI Images
Steps:
- Assign the gloabal variables
- Meta data Filtering
- Preprocessing and Spatial Gap filling
- Model Creation and Model Trainng
- Creating the predictions using the model and output comparison
import os
import warnings
warnings.filterwarnings("ignore")
os.environ['TF_ENABLE_ONEDNN_OPTS'] = '0'
os.environ['TF_CPP_MIN_LOG_LEVEL'] = '2'
os.environ["CUDA_VISIBLE_DEVICES"] = "1"
from glob import glob
from pathlib import Path
from libs.RASTERHELPER import RASTERHELPER as RH
from skimage.transform import resize
from matplotlib import pyplot as plt
import matplotlib.colors as mcolors
import pandas as pd
import numpy as np
import rasterio as rio
from libs.DLHELPER import DLHELPER as DH
import tensorflow as tf
tf.config.optimizer.set_jit(False)
from tensorflow.keras import models
from tensorflow.keras.models import Sequential, save_model
from tensorflow.keras.layers import ConvLSTM2D, BatchNormalization, Dense, Dropout, TimeDistributed, Conv3D
from tensorflow.keras.callbacks import EarlyStopping, CSVLogger, ModelCheckpoint, ReduceLROnPlateau
from tensorflow.keras.losses import MeanSquaredError
from tensorflow.keras.optimizers import Adam
from tensorflow.keras.metrics import RootMeanSquaredError
from datetime import datetime
import re
import random, shutil
RH=RH()
DH = DH()
STRIDE =8
N_EPOCH = 2
N_INPUTBANDS = 2
TARGET_SHAPE = (128, 128)
TARGET_SHAPE_X = (N_INPUTBANDS, 128, 128)
TARGET_SHAPE_Y = (128, 128)
TRAINING_BATCH_SIZE = 16
SEED = 50
MIN_R_H, MIN_R_W = 32, 32
PREDICTOR_NAMES = ["VV", "VH"]
PREDICTING_NAME = "NDVI"
DIR_MODEL_SAVE = './Models_S1S2/'
DIR_PLOTS = "./Plots_S1S2"
TRAIN_PATH = './../Data/AllCropS1S2_train_test/global_balanced_train.csv'
VAL_PATH = './../Data/AllCropS1S2_train_test/global_balanced_validation.csv'
TEST_PATH = './../Data/AllCropS1S2_train_test/global_balanced_test.csv'Preprocessing
- Filterout fileds which are smaller than 32 X 32 pixels
- Filterout images which has NaN pixels more then 10 % of their total pixels
- Filterout images which has incnsistency in metadata (i.e. Inconsistancy in Band Names, Image instance created but bands are not downloaded(happens due to broken connection during downloading))
print(f"\n\n{'='*40}\nLoading CSV Datasets\n{'='*40}")
TrainFilesDF = pd.read_csv(TRAIN_PATH).sample(frac=1).reset_index(drop=True)
ValidationFilesDF = pd.read_csv(VAL_PATH).sample(frac=1).reset_index(drop=True)
TestFilesDF = pd.read_csv(TEST_PATH).sample(frac=1).reset_index(drop=True)
print(f'\tTrain: {TrainFilesDF.shape}, Val: {ValidationFilesDF.shape}, Test: {TestFilesDF.shape}')
print(f"\n\n{'='*40}\nFilter out erroneous images\n{'='*40}")
TrainFilesDF = TrainFilesDF.copy()
ValidationFilesDF = ValidationFilesDF.copy()
results_train = TrainFilesDF.apply(RH.check_row_for_errors, axis=1, predictor_bands=PREDICTOR_NAMES, predicting_name=PREDICTING_NAME, nan_threshold=0.10, min_r_h=MIN_R_H, min_r_w=MIN_R_W, verbose=False)
TrainFilesDF[['IS_VALID', 'ERROR_MESSAGE', 'META_INFO']] = pd.DataFrame(results_train.tolist(), index=TrainFilesDF.index)
updated_train_df = TrainFilesDF[TrainFilesDF['IS_VALID']].reset_index(drop=True)
train_error_files = TrainFilesDF[~TrainFilesDF['IS_VALID']]
print(f"\tFiltering Train Done, Removed {len(train_error_files)} rows, Current Shape: {updated_train_df.shape}")
# Process Validation Data
print("\tFiltering Validation Data...")
results_validation = ValidationFilesDF.apply(RH.check_row_for_errors, axis=1, predictor_bands=PREDICTOR_NAMES, predicting_name=PREDICTING_NAME, nan_threshold=0.10, min_r_h=MIN_R_H,min_r_w=MIN_R_W,verbose=False)
ValidationFilesDF[['IS_VALID', 'ERROR_MESSAGE', 'META_INFO']] = pd.DataFrame(results_validation.tolist(), index=ValidationFilesDF.index)
updated_validation_df = ValidationFilesDF[ValidationFilesDF['IS_VALID']].reset_index(drop=True)
validation_error_files = ValidationFilesDF[~ValidationFilesDF['IS_VALID']]
print(f"\tFiltering Validation Done, Removed {len(validation_error_files)} rows, Current Shape: {updated_validation_df.shape}")
print(f"\n\n{'='*40}\nVisualising Random Train and Validation\n{'='*40}")
rand_index = min(random.choice(updated_train_df.index), random.choice(updated_validation_df.index))
rand_data_train = updated_train_df.iloc[rand_index]
rand_data_val = updated_validation_df.iloc[rand_index]
rand_train_s1_image, _, rand_train_s1_bnames = RH.load_geotiff(rand_data_train.FILEPATH_S1)
rand_train_s2_image, _, rand_train_s2_bnames = RH.load_geotiff(rand_data_train.FILEPATH_S2)
p_i = rand_train_s2_bnames.index(PREDICTING_NAME)
# RH.plot_from_bands(np.concatenate([rand_train_s1_image[0:2,:,:], rand_train_s2_image[p_i:p_i+1, :, :]], axis=0), PREDICTOR_NAMES + [PREDICTING_NAME], 'Raw Train: '+rand_data_train.FILEPATH_S1)
rand_val_s1_img, _, rand_val_s1_bnames = RH.load_geotiff(rand_data_val.FILEPATH_S1)
rand_val_s2_img, _, rand_val_s2_bnames = RH.load_geotiff(rand_data_val.FILEPATH_S2)
p_i = rand_val_s2_bnames.index(PREDICTING_NAME)
RH.plot_from_bands(np.concatenate([rand_val_s1_img[0:2, :, :], rand_val_s2_img[p_i:p_i+1, :, :]], axis=0), PREDICTOR_NAMES + [PREDICTING_NAME], 'Raw Val: '+rand_data_val.FILEPATH_S1)
========================================
Loading CSV Datasets
========================================
Train: (17, 18), Val: (9, 18), Test: (14, 18)
========================================
Filter out erroneous images
========================================
Filtering Train Done, Removed 5 rows, Current Shape: (12, 21)
Filtering Validation Data...
Filtering Validation Done, Removed 4 rows, Current Shape: (5, 21)
========================================
Visualising Random Train and Validation
========================================

The dataset size has smaller size. With ideal dataset it would look like the one below.
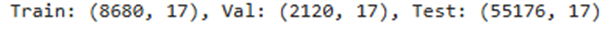
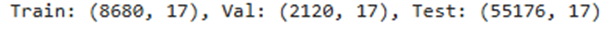
Feature Engineering
- Filling the image gaps which was created due to cloud masking or any other reason
- Normalizing the bands using min-max calling (Note the min max values for train test for future transfer learning)
- Creating New Features
- Radar Vegetation Index
- Band Cross Ratio
- Day of the Year
- Patching Images (Fields are of various sizes). But for the model the size needs to be consistant. Therefore, the images has been splitted into chips of
TARGET_SHAPE128x128 px
[ For filling the empty pixels NAVIER STOKES algorithm was used through opencv library. (https://www.math.ucla.edu/~bertozzi/papers/cvpr01.pdf) ]
print(f"\n\n{'='*40}\nLoading all images\n{'='*40}")
bands_train_predictor, bands_train_predicting, train_processing_error = RH.processFiles(dataframe=updated_train_df, predictor_names=PREDICTOR_NAMES, predicting_name=PREDICTING_NAME, filling_ksize = 1, plot_filling_comp = False, verbose = True)
print(f"\tFilling Train Done, Errors: {len(train_processing_error)}")
bands_validation_predictor, bands_validation_predicting, validation_processing_error = RH.processFiles( dataframe=updated_validation_df, predictor_names=PREDICTOR_NAMES, predicting_name=PREDICTING_NAME, filling_ksize = 1, plot_filling_comp = False, verbose = True)
print(f"\tFilling Validation Done, Errors: {len(validation_processing_error)}")
updated_train_df = RH.remove_error_indices(updated_train_df, train_processing_error)
updated_validation_df = RH.remove_error_indices(updated_validation_df, validation_processing_error)
print(f"\tCleaned train dataframe size: {updated_train_df.shape}")
print(f"\tCleaned validation dataframe size: {updated_validation_df.shape}")
print(f"\tGenerating fatures: Training")
bands_train_predictor = RH.feature_generation_rvi_bcr(bands_train_predictor)
print(f"\tGenerating Features: Validation")
bands_validation_predictor = RH.feature_generation_rvi_bcr(bands_validation_predictor)
bands_train_predictor, min_vals, max_vals = RH.normalize_bands_minmax(bands_train_predictor)
bands_validation_predictor, _, _ = RH.normalize_bands_minmax(bands_validation_predictor, min_vals=min_vals, max_vals=max_vals)
print("\tNormalization complete for both training and validation sets.")
print(f"\tExample normalized train image shape: {bands_train_predictor[0].shape}")
print(f"\tExample normalized validation image shape: {bands_validation_predictor[0].shape}")
print(f"\tMin value: {min_vals}(SAVE FOR FUTURE TRANSFER LEARNING) \n\tMax value:{max_vals}(SAVE FOR FUTURE TRANSFER LEARNING)")
print(f"\tAdding DOY band")
bands_train_predictor = RH.addDoyBand(bands_train_predictor, updated_train_df)
bands_validation_predictor = RH.addDoyBand(bands_validation_predictor, updated_validation_df)
RH.plot_distributions(updated_train_df, df_title = f'Train(Total: {updated_train_df.shape})', plot_save_dir=None)
RH.plot_distributions(updated_validation_df, df_title = f'Validation(Total: {updated_validation_df.shape}', plot_save_dir=None)
========================================
Loading all images
========================================
Filling Train Done, Errors: 0
Filling Validation Done, Errors: 0
Cleaned train dataframe size: (12, 21)
Cleaned validation dataframe size: (5, 21)
Generating fatures: Training
Generating Features: Validation
Normalization complete for both training and validation sets.
Example normalized train image shape: (4, 191, 164)
Example normalized validation image shape: (4, 142, 102)
Min value: [np.float32(-33.282623), np.float32(-40.947224), np.float32(0.009077747), np.float32(0.08207506)](SAVE FOR FUTURE TRANSFER LEARNING)
Max value:[np.float32(13.379636), np.float32(-2.1852489), np.float32(7.3922515), np.float32(879.58136)](SAVE FOR FUTURE TRANSFER LEARNING)
Adding DOY band
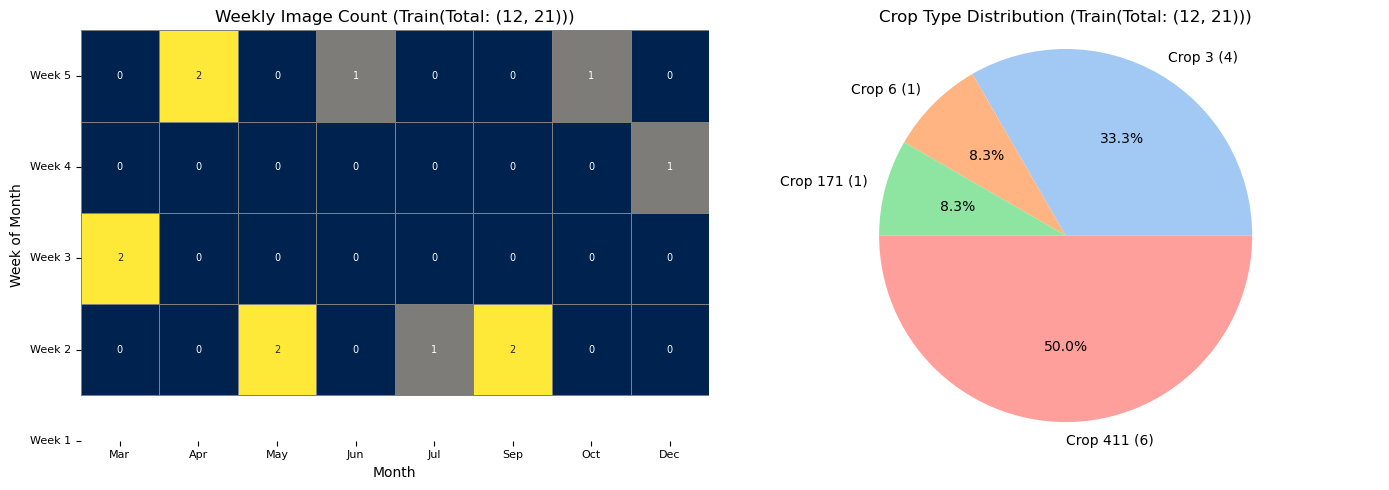
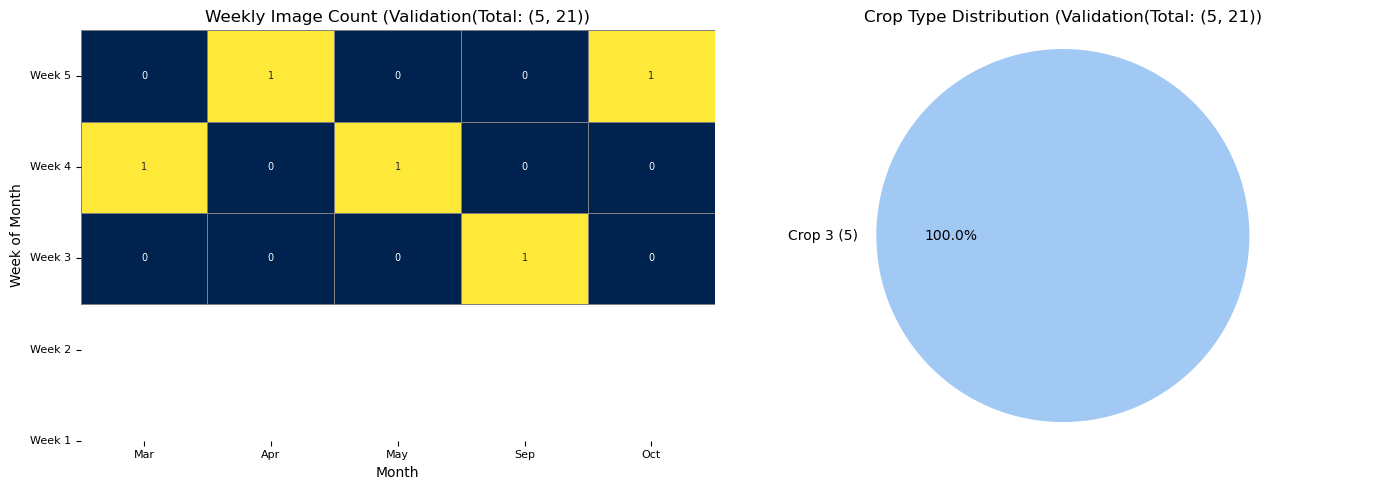
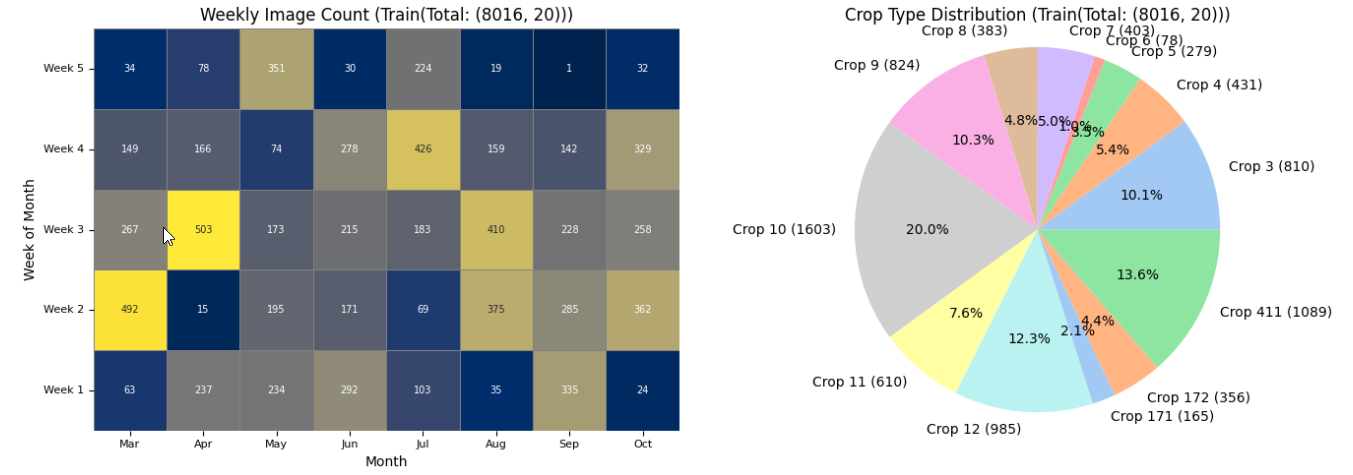
The plot has less number of images due to redued data size. With larger number of data it would look like the one below.
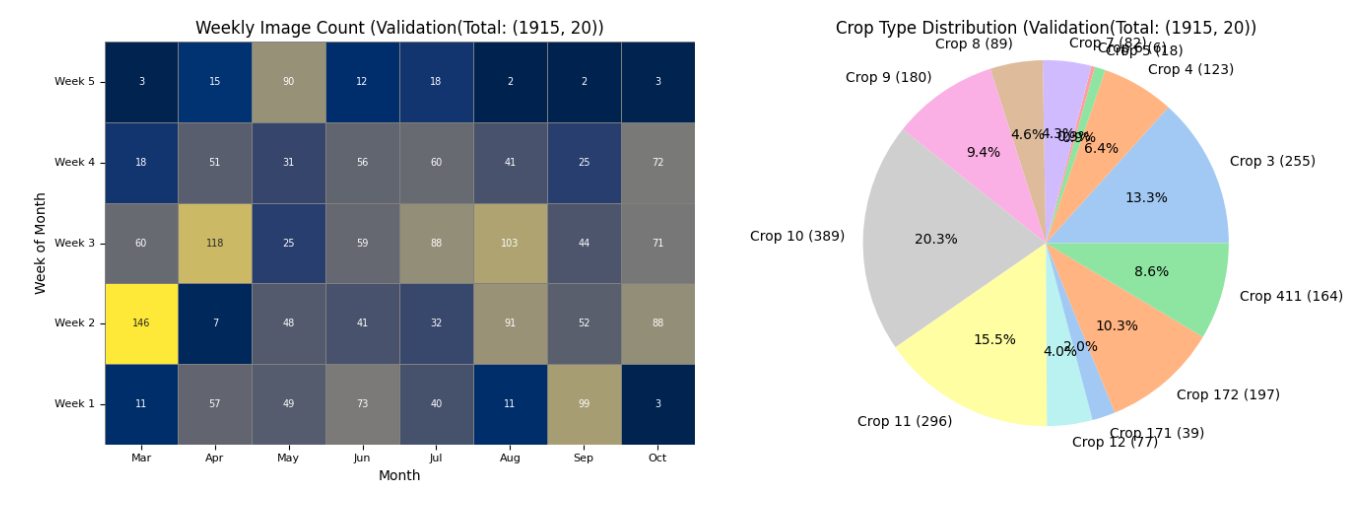
# Patch Extraction
print(f"\n\n{'='*40}\nPatching Images\n{'='*40}")
all_x_train_patches, all_y_train_patches, all_x_val_patches, all_y_val_patches = [], [], [], []
training_image_original_shapes = [] # Not used in this specific comparison, but good to keep
patch_size = TARGET_SHAPE
validation_image_original_shapes, all_x_val_patches_with_coords = [], [] # Store all validation patches WITH coords for reconstruction
for i, image_x in enumerate(bands_train_predictor):
image_y = bands_train_predicting[i]
original_unpadded_shape_x = image_x.shape
original_unpadded_shape_y = image_y.shape
x_patches_with_coords = RH.extract_patches_overlapping(image_array = image_x, patch_size = patch_size, stride = (TARGET_SHAPE[0]-STRIDE, TARGET_SHAPE[0]-STRIDE), pad_mode = 'symmetric') # or reflect
y_patches_with_coords = RH.extract_patches_overlapping(image_array = image_y, patch_size = patch_size, stride = (TARGET_SHAPE[0]-STRIDE, TARGET_SHAPE[0]-STRIDE), pad_mode = 'symmetric')
if len(x_patches_with_coords) != len(y_patches_with_coords):
print(f"Warning: Mismatch in number of patches for image {i}. Skipping.")
continue
for (x_patch, x_coords), (y_patch, y_coords) in zip(x_patches_with_coords, y_patches_with_coords):
all_x_train_patches.append(x_patch)
all_y_train_patches.append(y_patch)
if x_patches_with_coords:
last_x_coord = x_patches_with_coords[-1][1][1]
last_y_coord = x_patches_with_coords[-1][1][0]
padded_H = last_y_coord + patch_size[0]
padded_W = last_x_coord + patch_size[1]
num_channels_x = x_patches_with_coords[0][0].shape[0]
num_channels_y = y_patches_with_coords[0][0].shape[0]
training_image_original_shapes.append({'x_padded_shape': (num_channels_x, padded_H, padded_W),'x_unpadded_shape': original_unpadded_shape_x,'y_padded_shape': (num_channels_y, padded_H, padded_W),'y_unpadded_shape': original_unpadded_shape_y})
print(f"\tPatching Train Done")
for i, image_x in enumerate(bands_validation_predictor):
image_y = bands_validation_predicting[i]
original_unpadded_shape_x = image_x.shape
original_unpadded_shape_y = image_y.shape # e.g., (H, W)
x_patches_with_coords = RH.extract_patches_overlapping(image_array = image_x, patch_size = patch_size, stride = (TARGET_SHAPE[0]-STRIDE, TARGET_SHAPE[0]-STRIDE), pad_mode = 'symmetric')
# Ensure label images are (1, H, W) for patching consistency
y_patches_with_coords = RH.extract_patches_overlapping(image_array = image_y, patch_size = patch_size, stride = (TARGET_SHAPE[0]-STRIDE, TARGET_SHAPE[0]-STRIDE), pad_mode = 'symmetric')
if len(x_patches_with_coords) != len(y_patches_with_coords):
print(f"Warning: Mismatch in number of patches for validation image {i}. Skipping.")
continue
# Store individual patch data
for (x_patch, x_coords), (y_patch, y_coords) in zip(x_patches_with_coords, y_patches_with_coords):
all_x_val_patches.append(x_patch)
all_y_val_patches.append(y_patch)
# Store patches with their coordinates for later reconstruction
all_x_val_patches_with_coords.append((x_patch, x_coords))
if x_patches_with_coords:
last_x_coord = x_patches_with_coords[-1][1][1]
last_y_coord = x_patches_with_coords[-1][1][0]
padded_H = last_y_coord + patch_size[0]
padded_W = last_x_coord + patch_size[1]
num_channels_x = x_patches_with_coords[0][0].shape[0]
num_channels_y = y_patches_with_coords[0][0].shape[0] # Should be 1 for a mask
validation_image_original_shapes.append({ 'x_padded_shape': (num_channels_x, padded_H, padded_W), 'x_unpadded_shape': original_unpadded_shape_x, 'y_padded_shape': (num_channels_y, padded_H, padded_W),'y_unpadded_shape': original_unpadded_shape_y})
print(f"\tPatching Test Done")
print(f"\n\n{'='*40}\nReshaping\n{'='*40}")
X_train_patches = np.array(all_x_train_patches)
Y_train_patches = np.array(all_y_train_patches)
X_val_patches = np.array(all_x_val_patches)
Y_val_patches = np.array(all_y_val_patches)
print(f"\tArry Conversion Done")
# Transpose for Keras model input (N, H, W, C)
X_train_model_input = np.transpose(X_train_patches, (0, 2, 3, 1))
Y_train_model_input = np.transpose(Y_train_patches, (0, 2, 3, 1))
X_val_model_input = np.transpose(X_val_patches, (0, 2, 3, 1))
Y_val_model_input = np.transpose(Y_val_patches, (0, 2, 3, 1))
print(f"\tTransposing Done")========================================
Patching Images
========================================
Patching Train Done
Patching Test Done
========================================
Reshaping
========================================
Arry Conversion Done
Transposing Done
Model creation and training
We have experimented with U-Net network. The network is typically used for the segmentation tasks. However, with proper adaption, the architecture can also be adapted for the regression tasks. That is wahat has been done in this tutorial.
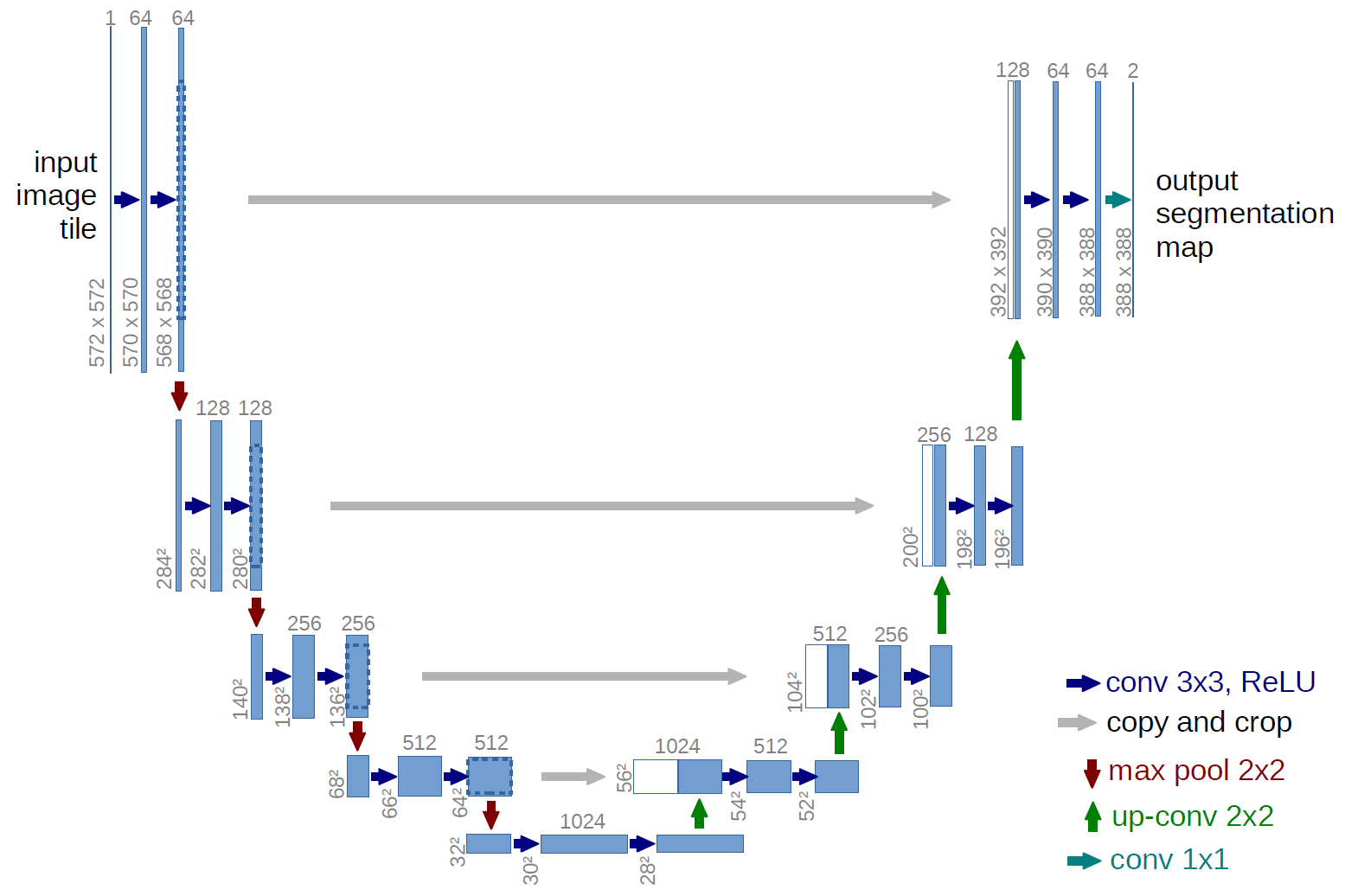
Credit: U-Net: Convolutional Networks for Biomedical Image Segmentation, Olaf Ronneberger , Philipp Fischer , Thomas Brox, Medical Image Computing and Computer-Assisted Intervention (MICCAI), Springer, LNCS, Vol.9351: 234--241, 2015, available at arXiv:1505.04597 [cs.CV]
- Model Creation
- Model Compilation(Optimizer, loss, Metrics)
- Defining callbacks(csv_logger, checkpoint, (early stopping not used))
- Model Training
- Model Saving
print(f"\n\n{'='*40}\nModel Creation\n{'='*40}")
print(f'\tShapes of the data:\n\tTrainX: {X_train_model_input.shape}, TrainY: {Y_train_model_input.shape}, \n \tValX: {X_val_model_input.shape}, ValY: {Y_val_model_input.shape}')
input_h, input_w, input_c = TARGET_SHAPE[0], TARGET_SHAPE[0], 5 # Assuming TARGET_SHAPE is (height, width)
model = DH.create_unet_model_with_dropout(input_height = input_h, input_width = input_w, num_input_bands = input_c, dropout_rate=0.0)
model.compile(optimizer=Adam(learning_rate=0.00001), loss='mae', metrics=['mae', 'mse', RootMeanSquaredError(name='rmse')])
os.makedirs(DIR_MODEL_SAVE, exist_ok=True)
training_timestamp = datetime.now().strftime("%Y-%m-%d_%H-%M")
log_file = os.path.join(DIR_MODEL_SAVE, f'Unet_TrainingLog_{training_timestamp}.csv')
model_file = os.path.join(DIR_MODEL_SAVE, f'Unet_Model_{training_timestamp}.keras')
lr_reducer = ReduceLROnPlateau(monitor='val_loss', factor=0.25, patience=10, mode='min', min_lr=1e-7, verbose=1)
csv_logger = CSVLogger(log_file, append=True)
early_stopping = EarlyStopping(monitor='val_loss', patience=25, restore_best_weights=True, verbose=1)
checkpoint = ModelCheckpoint(model_file, monitor='val_loss', save_best_only=True, verbose=1)
print(model.summary())
print(f"\n\n{'='*40}\nModel Traing\n{'='*40}")
if tf.config.list_physical_devices('GPU'):
print("\tTraining on GPU.")
with tf.device('GPU'):
history = model.fit(x=X_train_model_input, y=Y_train_model_input, batch_size=TRAINING_BATCH_SIZE, epochs=N_EPOCH, validation_data=(X_val_model_input, Y_val_model_input), callbacks=[csv_logger, checkpoint])
else:
print("\tNo GPU found, training on CPU.")
with tf.device('/CPU:0'):
history = model.fit(x=X_train_model_input, y=Y_train_model_input, batch_size=TRAINING_BATCH_SIZE, epochs=N_EPOCH, validation_data=(X_val_model_input, Y_val_model_input), callbacks=[csv_logger,checkpoint])
# Save the model after training
# model.save(model_file)
print(f"\tModel saved successfully.\n\tModel Path: {model_file}\n\tLog Path: {log_file}")
print(f"\n\n{'='*40}\nModel History\n{'='*40}")
history_df = pd.read_csv(log_file)
DH.plot_train_history(history_df, metrics=['loss', 'mae', 'mse', 'rmse'])========================================
Model Creation
========================================
Shapes of the data:
TrainX: (40, 128, 128, 5), TrainY: (40, 128, 128, 1),
ValX: (14, 128, 128, 5), ValY: (14, 128, 128, 1)
None
========================================
Model Traing
========================================
No GPU found, training on CPU.
Epoch 1/2
3/3 ━━━━━━━━━━━━━━━━━━━━ 0s 12s/step - loss: 1.1008 - mae: 1.1008 - mse: 1.6939 - rmse: 1.3015
Epoch 1: val_loss improved from None to 0.63899, saving model to ./Models_S1S2/Unet_Model_2025-09-10_14-46.keras
3/3 ━━━━━━━━━━━━━━━━━━━━ 59s 15s/step - loss: 1.0915 - mae: 1.0915 - mse: 1.6701 - rmse: 1.2923 - val_loss: 0.6390 - val_mae: 0.6390 - val_mse: 0.4539 - val_rmse: 0.6737
Epoch 2/2
3/3 ━━━━━━━━━━━━━━━━━━━━ 0s 12s/step - loss: 1.0538 - mae: 1.0538 - mse: 1.5894 - rmse: 1.2607
Epoch 2: val_loss did not improve from 0.63899
3/3 ━━━━━━━━━━━━━━━━━━━━ 43s 14s/step - loss: 1.0613 - mae: 1.0613 - mse: 1.6097 - rmse: 1.2687 - val_loss: 0.6484 - val_mae: 0.6484 - val_mse: 0.4662 - val_rmse: 0.6828
Model saved successfully.
Model Path: ./Models_S1S2/Unet_Model_2025-09-10_14-46.keras
Log Path: ./Models_S1S2/Unet_TrainingLog_2025-09-10_14-46.csv
========================================
Model History
========================================
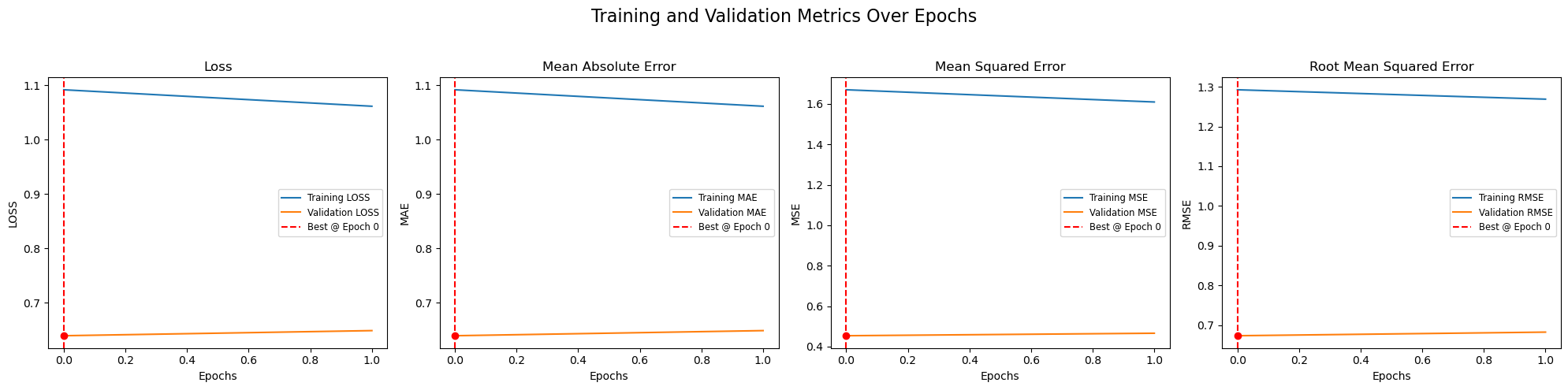
The plot has less number of images and epochs due to redued data size. With larger number of data it would look like the one below.
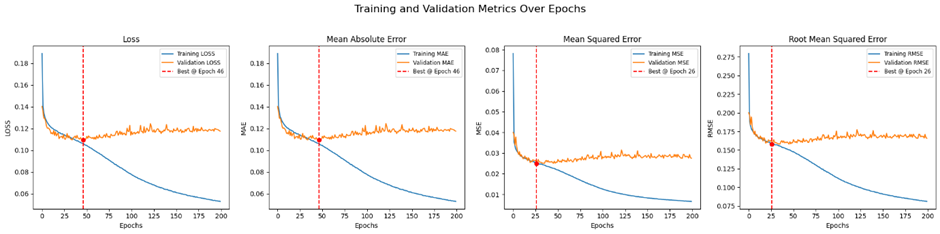
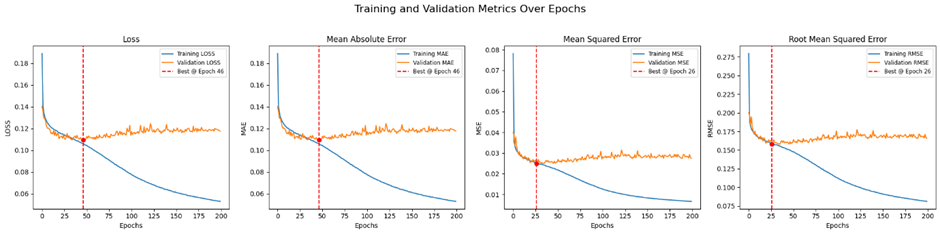
Model performance
- Make the predictions for the splitted chips
- Reconstruct the predicted image from chips
- Compare the reconstructed predicted NDVI with the ground truth
print(f"\n\n{'='*40}\nMaking predictions on validation patches\n{'='*40}")
predicted_val_patches_model_output = model.predict(X_val_model_input) # Shape (num_patches, H, W, 1)
predicted_val_patches_for_reconstruction = np.transpose(predicted_val_patches_model_output, (0, 3, 1, 2))
print(f"\tShape of predicted_val_patches_for_reconstruction: {predicted_val_patches_for_reconstruction.shape}")
print(f"\n\n{'='*40}\nReconstructing full predicted and actual images\n{'='*40}")
reconstructed_predicted_images, reconstructed_actual_images, reconstructed_vv_images, reconstructed_vh_images = [],[],[],[],
patch_count_per_image = []
current_patch_idx = 0
# Re-extracting patches with coords for each original full image (X and Y), This allows us to map predicted patches back to their original image context for reconstruction
original_val_image_x_patches_with_coords_list = []
original_val_image_y_patches_with_coords_list = []
for i, full_x_image in enumerate(bands_validation_predictor):
full_y_image = bands_validation_predicting[i]
x_patches_of_this_image = RH.extract_patches_overlapping(image_array = full_x_image, patch_size = patch_size, stride = (TARGET_SHAPE[0]-STRIDE, TARGET_SHAPE[0]-STRIDE), pad_mode = 'symmetric')
y_patches_of_this_image = RH.extract_patches_overlapping(image_array = full_y_image, patch_size = patch_size,stride = (TARGET_SHAPE[0]-STRIDE, TARGET_SHAPE[0]-STRIDE), pad_mode = 'symmetric')
original_val_image_x_patches_with_coords_list.append(x_patches_of_this_image)
original_val_image_y_patches_with_coords_list.append(y_patches_of_this_image)
current_global_patch_idx = 0
for img_idx, shape_info in enumerate(validation_image_original_shapes):
# Get the original full image and label for reference (VV/VH bands)
original_full_x_image = bands_validation_predictor[img_idx]
original_full_y_image = bands_validation_predicting[img_idx] # This is your original label mask
num_patches_in_this_image = len(original_val_image_x_patches_with_coords_list[img_idx])
predicted_patches_for_this_image_raw = predicted_val_patches_for_reconstruction[current_global_patch_idx : current_global_patch_idx + num_patches_in_this_image]
# Create (patch_data, (y,x)) tuples for reconstruction function
predicted_patches_with_coords_for_reconstruction = []
for j in range(num_patches_in_this_image):
original_coords = original_val_image_x_patches_with_coords_list[img_idx][j][1]
predicted_patch_data = predicted_patches_for_this_image_raw[j]
predicted_patches_with_coords_for_reconstruction.append((predicted_patch_data, original_coords))
# Reconstruct the predicted NDVI image
reconstructed_pred_ndvi = RH.reconstruct_overlapping(predicted_patches_with_coords = predicted_patches_with_coords_for_reconstruction, original_padded_shape=shape_info['y_padded_shape'], original_unpadded_shape=shape_info['y_unpadded_shape'], patch_size = TARGET_SHAPE)
reconstructed_predicted_images.append(reconstructed_pred_ndvi.squeeze(axis=0)) # Squeeze channel for 2D
reconstructed_actual_images.append(original_full_y_image) # This assumes original_full_y_image is (H,W)
reconstructed_vv_images.append(original_full_x_image[0, :, :]) # Assuming VV is band 0
reconstructed_vh_images.append(original_full_x_image[1, :, :]) # Assuming VH is band 1
current_global_patch_idx += num_patches_in_this_image
print(f"\tNumber of reconstructed predicted images: {len(reconstructed_predicted_images)}, Actual Images: {len(reconstructed_actual_images)}")
print(f"\tShape of first reconstructed shape predicted image: {reconstructed_predicted_images[1].shape} actual: {reconstructed_actual_images[1].shape}")
print(f"\tPreparing labels for validation predictions")
val_dates = updated_validation_df['DATE'].dt.strftime('%Y-%m-%d')
val_field_nums = updated_validation_df['FIELDNUM'].astype(str)
val_crop_types = updated_validation_df['CROPTYPE'].astype(str)
val_crop_groups = updated_validation_df['CROPGROUP'].astype(str)
# Filter val_labels to match the number of reconstructed images (which should be the number of original images)
val_labels_for_reconstructed_images = ("Field_" + val_field_nums + "_" + val_crop_groups + "_" + val_crop_types + "_" + val_dates ).tolist()
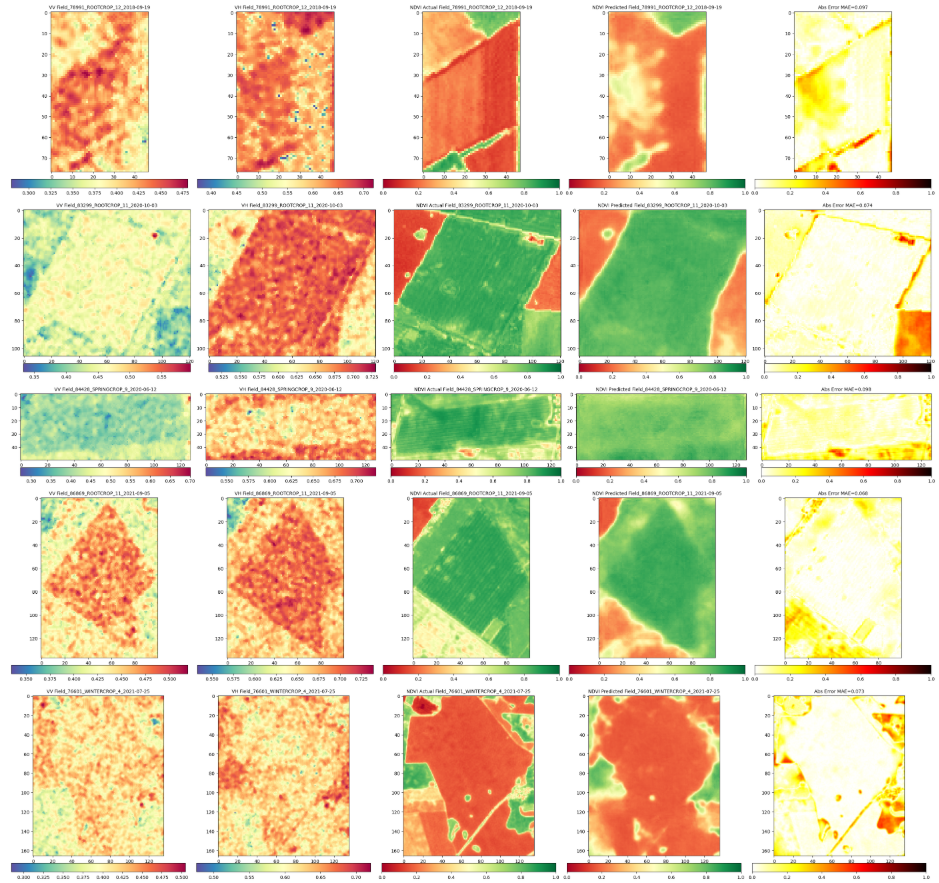
print(f"\n\n{'='*40}\nFieldwise Comparison of Predicted and round Truth NDVI\n{'='*40}")
DIR_PLOTS = f'{DIR_PLOTS}/Reconstructed_Val_Comp_Unet_{training_timestamp}' # New directory for reconstructed plots
c_reconstructed = 2 #len(reconstructed_predicted_images) # Compare all reconstructed images
print(f"\tCalling comparison function for {c_reconstructed} reconstructed images")
RH.compareNDVIs_with_VV_VH(actual_ndvis = reconstructed_actual_images[:c_reconstructed], predicted_ndvis = reconstructed_predicted_images[:c_reconstructed], actual_vv = reconstructed_vv_images[:c_reconstructed],
actual_vh = reconstructed_vh_images[:c_reconstructed], dates = val_labels_for_reconstructed_images[:c_reconstructed], export_path = DIR_PLOTS, export_plot = False) # Set to True to save plots
print(f"\tComparison of reconstructed images complete. Check '{DIR_PLOTS}' for plots.")========================================
Making predictions on validation patches
========================================
1/1 ━━━━━━━━━━━━━━━━━━━━ 3s 3s/step
Shape of predicted_val_patches_for_reconstruction: (14, 1, 128, 128)
========================================
Reconstructing full predicted and actual images
========================================
Number of reconstructed predicted images: 5, Actual Images: 5
Shape of first reconstructed shape predicted image: (103, 209) actual: (1, 103, 209)
Preparing labels for validation predictions
========================================
Fieldwise Comparison of Predicted and round Truth NDVI
========================================
Calling comparison function for 2 reconstructed images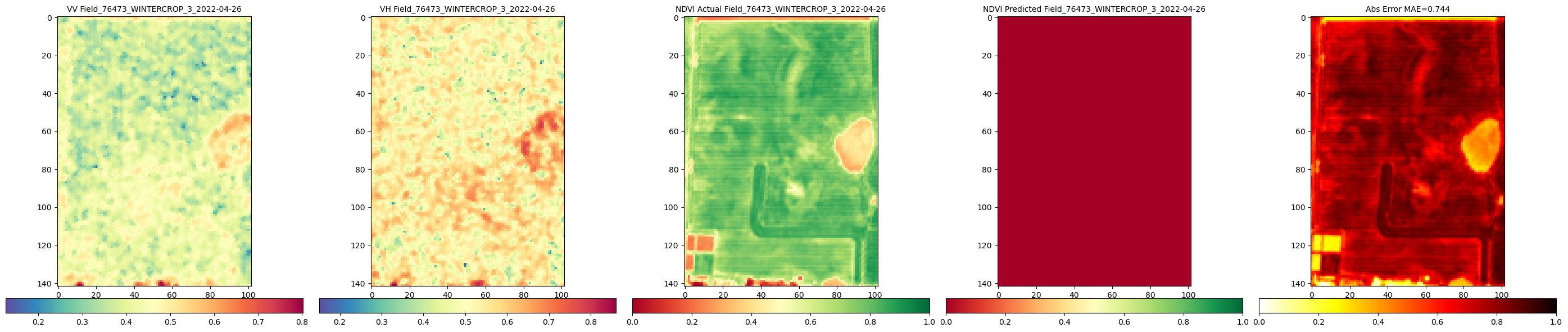
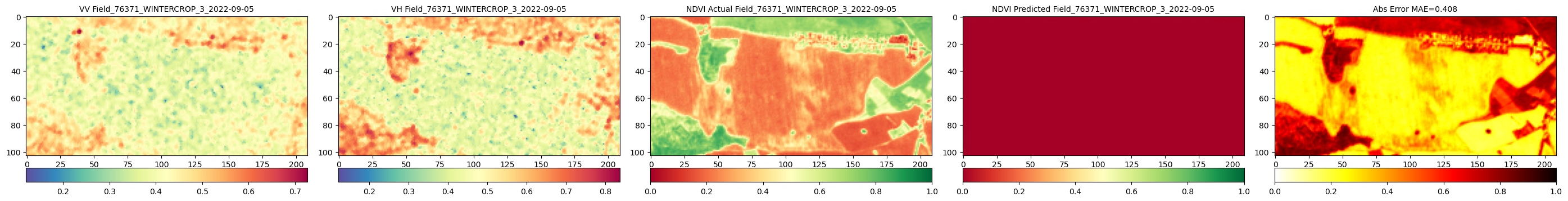
Comparison of reconstructed images complete. Check './Plots_S1S2/Reconstructed_Val_Comp_Unet_2025-09-10_14-46' for plots.The plot has less number of images due to redued data size. With larger number of data and training more epochs it would look like the one below.