Section 8
Time Series Mapping of Groundwater Levels
Estimated Time: ~30 minutes
Now let’s use the trained models to generate groundwater-level predictions across the study area. Raster datasets will be imported to visualize spatial patterns in the predicted groundwater dynamics. Time-indexed rasters will enable temporal analyses and animations to track changes across seasons and years. All layers will be harmonized in projection, resolution, and extent to ensure accurate spatiotemporal comparisons.
Define target year, file paths, and shapefile
base_path <- "C:/Users/raza"
target_year <- "2002"
folder_paths <- c(
"C:/Users/raza/project/ET",
"C:/Users/raza/project/EVI",
"C:/Users/raza/project/NDVI",
"C:/Users/raza/project/Prec",
"C:/Users/raza/project/Temp"
)
# Load reference raster for projection and alignment
reference_raster <- raster("C:/Users/raza/project/ASP.tif")
reference_coordinates <- coordinates(reference_raster)
# Load and reproject shapefile
shapefile <- readOGR("C:/Users/raza/project/borders/Brandenburg.shp")
shapefile <- spTransform(shapefile, CRS("+proj=utm +zone=32 +datum=WGS84 +units=m +no_defs"))
# Initialize empty dataframe for results
final_data <- data.frame(Date = as.Date(character(0)), Value = numeric(0), X = numeric(0), Y = numeric(0))
Loop through raster folders and extract values
extract_layer_data <- function(folder_path, target_year, reference_raster, shapefile) {
folder_data <- data.frame()
files <- list.files(folder_path, pattern = "\\.tif$", full.names = TRUE)
for (month in 1:12) {
matching_files <- grep(paste0(target_year, ".", sprintf("%02d", month)), files, value = TRUE)
for (file_path in matching_files) {
raster_layer <- raster(file_path)
raster_layer <- projectRaster(raster_layer, crs = crs(reference_raster))
raster_layer <- crop(raster_layer, extent(shapefile))
raster_layer <- mask(raster_layer, shapefile)
values <- extract(raster_layer, reference_coordinates)
coords <- coordinates(reference_raster)
date <- as.Date(paste(target_year, sprintf("%02d", month), "01", sep = "-"))
variable_name <- substr(basename(file_path), 1, 3)
new_row <- data.frame(Date = date, Value = values, X = coords[,1], Y = coords[,2])
colnames(new_row)[colnames(new_row) == "Value"] <- variable_name
folder_data <- bind_rows(folder_data, new_row)
}
}
return(folder_data)
}
# Loop through all folders and combine
for (folder in folder_paths) {
folder_data <- extract_layer_data(folder, target_year, reference_raster, shapefile)
final_data <- bind_rows(final_data, folder_data)
}
Reshape and clean data
# Pivot to wide format based on raster variable names
final_data_wide <- final_data %>%
group_by(Date, X, Y) %>%
summarise_all(list(~ if(all(is.na(.))) NA else na.omit(.))) %>%
ungroup()
final_data_wide <- na.omit(final_data_wide)
# Save wide data for predictions
write.csv(final_data_wide, "C:/Users/raza/project/data_for_pred_2022.csv", row.names = FALSE)
Extract raster values for additional layers
raster_dir <- "C:/Users/raza/project/brandenburg"
raster_files <- list.files(raster_dir, pattern = "\\.tif$", full.names = TRUE)
csv_data <- read.csv("C:/Users/raza/project/data_for_pred_2022.csv")
extract_raster_values <- function(raster_file, x, y) {
raster_layer <- raster(raster_file)
extract(raster_layer, cbind(x, y))
}
for (raster_file in raster_files) {
raster_name <- sub(".tif", "", basename(raster_file))
csv_data[[raster_name]] <- extract_raster_values(raster_file, csv_data$X, csv_data$Y)
}
# Reorder and scale columns
desired_order <- c("Date", "X", "Y", "SLOPE", "ASP", "DEM", "LNF", "TPI", "TWI", "TRI", "PREC", "TEMP", "NDVI", "ET", "EVI")
csv_data <- csv_data[, desired_order]
csv_data$NDVI <- csv_data$NDVI / 10000
# Convert coordinates to WGS84
coordinates(csv_data) <- ~ X + Y
proj4string(csv_data) <- CRS("+proj=utm +zone=32 +datum=WGS84 +units=m +no_defs")
csv_data <- spTransform(csv_data, CRS("+proj=longlat +datum=WGS84"))
csv_data$Lon <- coordinates(csv_data)[,1]
csv_data$Lat <- coordinates(csv_data)[,2]
# Save final CSV
write.csv(csv_data, "C:/Users/raza/project/b_data_for_prediction/data_for_pred_2022_v2.csv", row.names = FALSE)
Create Monthly Prediction Rasters using Random Forest Model
library(raster)
library(sp)
library(dplyr)
# -----------------------------
# Define directories and paths
# -----------------------------
model_dir <- file.path(base_path, "project/models")
test_data_dir <- file.path(base_path, "project", "b_data_for_prediction")
output_dir <- file.path(base_path, "project/plot")
reference_raster_path <- file.path(base_path, "project/brandenburg", "DEM.tif")
# -----------------------------
# Prepare reference raster
# -----------------------------
reference_raster <- raster(reference_raster_path)
projected_raster <- projectRaster(reference_raster, crs = CRS("+proj=longlat +datum=WGS84"))
template_raster <- raster(projected_raster)
# -----------------------------
# Load trained Random Forest model
# -----------------------------
model_file <- file.path(model_dir, "RF_2001-05.rda")
load(model_file)
# -----------------------------
# Process data year by year
# -----------------------------
for (year in 2001:2005) {
test_data_file <- file.path(test_data_dir, paste0("data_for_pred_", year, "_v2.csv"))
if (!file.exists(test_data_file)) {
message(paste("File not found:", test_data_file))
next
}
test_data <- read.csv(test_data_file) %>% na.omit()
if (nrow(test_data) == 0) {
message(paste("No valid data for year:", year))
next
}
# Predict groundwater or variable of interest
preds <- predict(rf_model, data = test_data)
test_data$Predictions <- preds$predictions
# Ensure Date is in proper format
test_data$Date <- as.Date(test_data$Date)
unique_months <- sort(unique(format(test_data$Date, "%Y-%m")))
# -----------------------------
# Create monthly rasters
# -----------------------------
for (month in unique_months) {
month_data <- test_data %>%
filter(format(Date, "%Y-%m") == month)
if (nrow(month_data) == 0) next
# Convert to spatial points
sp_data <- month_data %>%
select(Lon, Lat, Predictions)
coordinates(sp_data) <- c("Lon", "Lat")
# Rasterize
monthly_raster <- rasterize(sp_data, template_raster, field = "Predictions", fun = mean)
# Define output path
output_file <- file.path(output_dir, paste0("monthly_rf_", month, ".tif"))
# Write raster
writeRaster(monthly_raster, filename = output_file, format = "GTiff", overwrite = TRUE)
}
message(paste("Completed processing for year:", year))
}
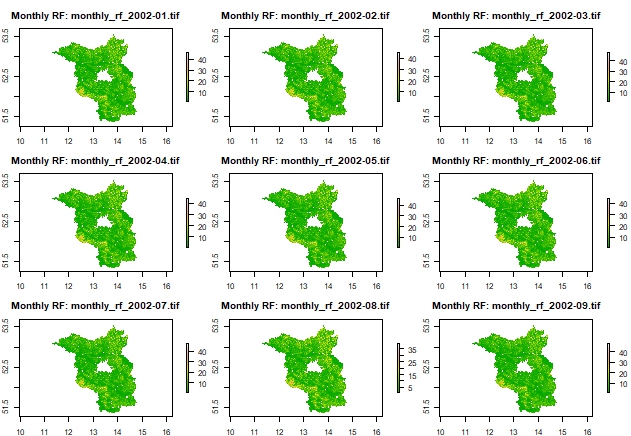
Plot 3x3 grid of monthly rasters as examples
# Get list of created rasters
raster_files <- list.files(output_dir, pattern = "monthly_rf_.*\\.tif$", full.names = TRUE)
# Check and plot up to 9 rasters
if (length(raster_files) > 0) {
num_to_plot <- min(9, length(raster_files))
selected_files <- raster_files[1:num_to_plot]
# Set plotting layout
par(mfrow = c(3, 3), mar = c(2, 2, 3, 4))
for (i in seq_along(selected_files)) {
r <- raster(selected_files[i])
plot(r, main = paste0("Monthly RF: ", basename(selected_files[i])),
col = terrain.colors(100))
}
par(mfrow = c(1, 1)) # Reset layout
} else {
message("No raster files found for plotting.")
}
Create monthly rasters using RFsp model
# Load libraries
library(doParallel)
library(landmap)
library(raster)
library(data.table)
library(dplyr)
library(plyr)
# Define directories
model_dir <- file.path(base_path, "project/models")
test_data_dir <- file.path(base_path, "project", "b_data_for_prediction")
output_dir <- file.path(base_path, "project/plot")
reference_raster_path <- file.path(base_path, "project/brandenburg", "DEM.tif")
# Load RFsp model
model_file <- file.path(model_dir, "RFsp_2001-05.rda")
load(model_file)
# Load reference raster and project to WGS84
reference_raster <- raster(reference_raster_path)
new_crs <- CRS("+proj=longlat +datum=WGS84")
reference_raster <- projectRaster(reference_raster, crs = new_crs)
# Function to create monthly rasters for a given year
process_year <- function(year) {
message("Processing year: ", year)
# Load test data
test_data_file <- file.path(test_data_dir, paste0("data_for_pred_1", year, "_v2.csv"))
if (!file.exists(test_data_file)) {
warning("File not found: ", test_data_file)
return(NULL)
}
test_data <- fread(test_data_file) %>% na.omit()
test_data[, staid := .I]
# Define bounding box and create fishnet grid
bbox <- extent(min(test_data$Lon), max(test_data$Lon), min(test_data$Lat), max(test_data$Lat))
bbox <- as(bbox, "SpatialPolygons")
proj4string(bbox) <- new_crs
fishnet <- raster(bbox)
res(fishnet) <- c(0.01, 0.01)
fishnet[] <- runif(ncell(fishnet))
fishnet <- as(fishnet, "SpatialPixelsDataFrame")
# Prepare date list
unique_dates <- unique(test_data$Date)
final_results <- list()
for (date in unique_dates) {
message(" Processing date: ", date)
# Filter and prepare spatial data
date_data <- test_data[Date == date]
spatial_points <- SpatialPointsDataFrame(coords = date_data[, .(Lon, Lat)], data = date_data, proj4string = new_crs)
# Compute buffer distances
dist_map <- landmap::buffer.dist(spatial_points["staid"], fishnet[1], as.factor(1:nrow(spatial_points)))
# Join distance info with test data
merged_data <- cbind(date_data, over(spatial_points, dist_map))
merged_data <- plyr::join(date_data, merged_data, by = "staid")
# Select and reorder columns
vars <- c("Date", "Lat", "Lon", "SLOPE", "ASP", "DEM", "LNF", "TPI", "TWI", "TRI", "PREC", "TEMP", "NDVI", "ET", "EVI")
other_vars <- setdiff(names(merged_data), vars)
merged_data <- merged_data[, c(vars, other_vars), with = FALSE]
merged_data <- merged_data[, !names(merged_data) %in% c("staid", "Lon.1", "Lat.1"), with = FALSE]
merged_data <- na.omit(merged_data)
# Predict with RFsp
preds <- predict(rfsp_model, data = merged_data)
merged_data$Predictions <- preds$predictions
final_results[[as.character(date)]] <- merged_data
}
# Combine results for the year
combined_results <- rbindlist(final_results)
# Save CSV output
output_csv <- file.path(output_dir, paste0(year, "_pred_rfsp1.csv"))
fwrite(combined_results[, .(Date, Lat, Lon, Predictions)], output_csv)
# Create monthly rasters
combined_results[, Date := as.Date(Date)]
months <- unique(format(combined_results$Date, "%Y-%m"))
for (month in months) {
message(" Creating raster for month: ", month)
monthly_data <- combined_results[format(Date, "%Y-%m") == month]
sp_data <- SpatialPointsDataFrame(
coords = monthly_data[, .(Lon, Lat)],
data = monthly_data[, .(Predictions)],
proj4string = new_crs
)
monthly_raster <- rasterize(sp_data, reference_raster, field = "Predictions")
output_file <- file.path(output_dir, paste0("monthly_rfsp_", month, ".tif"))
writeRaster(monthly_raster, filename = output_file, format = "GTiff", overwrite = TRUE)
}
message("Completed year: ", year)
}
# Run processing loop
for (year in 2001:2005) {
process_year(year)
}
message("All years processed successfully.")
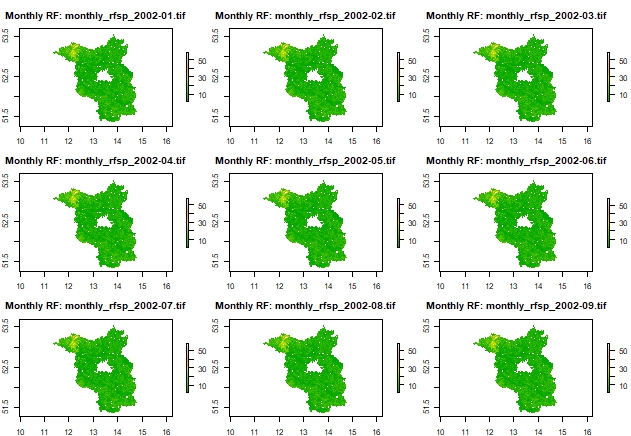
Plot 3x3 grid of monthly rasters as examples
# Get list of created rasters
raster_files <- list.files(output_dir, pattern = "monthly_rfsp_.*\\.tif$", full.names = TRUE)
# Check and plot up to 9 rasters
if (length(raster_files) > 0) {
num_to_plot <- min(9, length(raster_files))
selected_files <- raster_files[1:num_to_plot]
# Set plotting layout
par(mfrow = c(3, 3), mar = c(2, 2, 3, 4))
for (i in seq_along(selected_files)) {
r <- raster(selected_files[i])
plot(r, main = paste0("Monthly RF: ", basename(selected_files[i])),
col = terrain.colors(100))
}
par(mfrow = c(1, 1)) # Reset layout
} else {
message("No raster files found for plotting.")
}
Create predictions with RFSI model
# Load required libraries
library(sp)
library(sf)
library(raster)
library(meteo)
library(data.table)
library(dplyr)
# Load input data and model
load("stfdf_2001-05.rda")
load("../models/RFSI_2001-05.rda")
# Define covariates
covariates <- c("GWD", "Lat", "Lon", "SLOPE", "ASP", "DEM", "LNF", "TPI", "TWI", "TRI",
"PREC", "TEMP", "NDVI", "ET", "EVI")
# Extract and clean data from STFDF
GWD_df <- as.data.frame(stfdf)[, c("sp.ID", "time", covariates)]
GWD_df <- GWD_df[complete.cases(GWD_df), ]
# Prepare temporal information
time <- sort(unique(GWD_df$time))
days <- gsub("-", "", time, fixed = TRUE)
# Convert to clean dataframe for later use
newdata <- GWD_df %>% na.omit()
# Define years for processing
years_of_interest <- 2001:2005
# Loop through each year
for (year in years_of_interest) {
message("Processing year: ", year)
# Load test data
test_data_file <- file.path(base_path, "project", "b_data_for_prediction",
paste0("data_for_pred_", year, "_v2.csv"))
if (!file.exists(test_data_file)) {
warning("Test data missing for ", year)
next
}
data <- fread(test_data_file) %>% na.omit()
# Convert test data to spatial object
spatial_points_df <- SpatialPointsDataFrame(coords = data[, .(Lon, Lat)], data = data,
proj4string = CRS("+proj=longlat +datum=WGS84"))
# Order columns and remove redundant fields
desired_order <- c("Date", "Lat", "Lon", "SLOPE", "ASP", "DEM", "LNF", "TPI", "TWI",
"TRI", "PREC", "TEMP", "NDVI", "ET", "EVI")
rest <- setdiff(names(data), desired_order)
data <- data[, c(desired_order, rest), with = FALSE]
data <- data[, !names(data) %in% c("staid", "Lon.1", "Lat.1", "X", "Y", "optional"), with = FALSE]
data <- na.omit(data)
# Convert data to sf
loc_sf <- st_as_sf(data, coords = c("Lon", "Lat"), crs = 4326)
newdata_sf <- st_as_sf(newdata, coords = c("Lon", "Lat"), crs = 4326)
loc_sf$Date <- as.Date(loc_sf$Date)
newdata_sf$time <- as.Date(newdata_sf$time)
# Compute nearest observations
unique_dates <- unique(loc_sf$Date)
nearest_obs_list <- list()
for (current_date in unique_dates) {
message(" Finding nearest observations for date: ", current_date)
loc_day <- loc_sf[loc_sf$Date == current_date, ]
newdata_day <- newdata_sf[newdata_sf$time == current_date, ]
if (nrow(loc_day) == 0 || nrow(newdata_day) == 0) next
nearest_obs <- meteo::near.obs(
locations = loc_day,
observations = newdata_day,
obs.col = "GWD",
n.obs = 100
)
nearest_obs_list[[as.character(current_date)]] <- nearest_obs
}
# Combine nearest observations
nearest_obs_combined <- do.call(rbind, nearest_obs_list)
nearest_obs_combined$date <- gsub("\\.\\d+$", "", rownames(nearest_obs_combined))
rownames(nearest_obs_combined) <- NULL
nearest_obs_combined <- nearest_obs_combined[, c("date", setdiff(names(nearest_obs_combined), "date"))]
# Merge with spatial data
merged_data <- cbind(st_drop_geometry(loc_sf), nearest_obs_combined)
# Prepare for prediction
merged_data <- merged_data[, !names(merged_data) %in% c("geometry", "Date")]
# Predict using RFSI model
message(" Predicting groundwater depth for ", year)
predictions <- predict(rfsi_model, merged_data)
loc_sf$Predictions <- predictions$prediction
# Convert to dataframe for rasterization
loc_df <- st_drop_geometry(loc_sf)
coords <- st_coordinates(loc_sf)
loc_df <- cbind(coords, loc_df)
colnames(loc_df)[1:2] <- c("Lon", "Lat")
# Save predictions as CSV
output_csv <- file.path(base_path, "project", paste0(year, "_pred_rfsi.csv"))
fwrite(loc_df, output_csv)
# Load and project reference raster
reference_raster <- raster(file.path(base_path, "project/brandenburg", "DEM.tif"))
projected_raster <- projectRaster(reference_raster, crs = CRS("+proj=longlat +datum=WGS84"))
# Create monthly rasters
loc_df$Date <- as.Date(loc_df$Date)
months <- unique(format(loc_df$Date, "%Y-%m"))
output_dir <- file.path(base_path, "plot", "del")
for (month in months) {
message(" Creating raster for month: ", month)
monthly_data <- loc_df[format(loc_df$Date, "%Y-%m") == month, ]
sp_data <- SpatialPointsDataFrame(coords = monthly_data[, c("Lon", "Lat")],
data = data.frame(Predictions = monthly_data$Predictions),
proj4string = CRS("+proj=longlat +datum=WGS84"))
monthly_raster <- rasterize(sp_data, projected_raster, field = "Predictions")
output_file <- file.path(output_dir, paste0("monthly_rfsi_", month, ".tif"))
writeRaster(monthly_raster, filename = output_file, format = "GTiff", overwrite = TRUE)
}
message("Completed year: ", year)
}
message("All RFSI predictions completed successfully.")
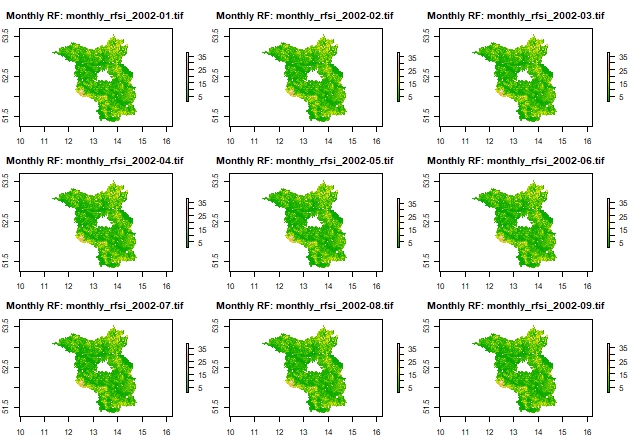
Plot 3x3 grid of monthly rasters as examples
# Get list of created rasters
raster_files <- list.files(output_dir, pattern = "monthly_rfsi_.*\\.tif$", full.names = TRUE)
# Check and plot up to 9 rasters
if (length(raster_files) > 0) {
num_to_plot <- min(9, length(raster_files))
selected_files <- raster_files[1:num_to_plot]
# Set plotting layout
par(mfrow = c(3, 3), mar = c(2, 2, 3, 4))
for (i in seq_along(selected_files)) {
r <- raster(selected_files[i])
plot(r, main = paste0("Monthly RF: ", basename(selected_files[i])),
col = terrain.colors(100))
}
par(mfrow = c(1, 1)) # Reset layout
} else {
message("No raster files found for plotting.")
}