Section 2
Curate and harmonize groundwater time series; identify and address quality issues
Estimated Time: ~1hr
In this section, we explore and prepare the groundwater monitoring data set. We then conduct a data quality assessment to determine the total number of wells to be used in this study to exclude anthropogenic and extreme climate conditions. Several criteria were applied to this initial selection process: (1) the consistency of data at each well; (2) the observational uncertainty related to the head values in the well database (RMSE =< 0.25m); and (3) continuous functionality of the well.
Please make sure that you have installed all the packages mentioned in the previous sections.
Curate and harmonize groundwater time series; identify and address quality issues
Identify CSV files starting from year 2000 and optionally remove others
folder_path <- "C:/Users/raza/project/ObsWell_based_data"
csv_files <- list.files(path = folder_path, pattern = "\\.csv$", full.names = TRUE)
num_files_meeting_condition <- 0
for (csv_file in csv_files) {
data <- read.csv(csv_file)
first_date <- dmy(data$Date[1])
if (!is.na(first_date) && year(first_date) >= 2000) {
num_files_meeting_condition <- num_files_meeting_condition + 1
cat("File:", basename(csv_file), "\n")
print(head(data$Date, 5))
# Remove file if needed
# unlink(csv_file)
}
}
cat("Number of files meeting condition:", num_files_meeting_condition, "\n")
Extract yearly data and add well ID
source_folder <- folder_path
destination_folder <- "C:/Users/raza/project/monthly_data"
dir_create(destination_folder)
specific_years <- 2001:2005
filter_and_add_filename <- function(data, year, filename) {
data <- data %>%
mutate(Date = dmy(Date)) %>%
filter(year(Date) == year) %>%
mutate(ID = filename)
return(data)
}
csv_files <- dir_ls(source_folder, regexp = "\\.csv$")
for (yr in specific_years) {
data_list <- list()
for (csv_file in csv_files) {
data <- read.csv(csv_file, stringsAsFactors = FALSE)
filename <- substr(tools::file_path_sans_ext(basename(csv_file)), 1, 8)
data_list[[filename]] <- filter_and_add_filename(data, yr, filename)
}
combined_data <- bind_rows(data_list)
write.csv(combined_data, paste0(destination_folder, "/", yr, ".csv"), row.names = FALSE)
cat("Year", yr, "data saved.\n")
}
Merge coordinates by Well ID
coordinate_file <- "C:/Users/raza/project/example/stammdaten_2023_07.csv"
coordinates_data <- read.csv(coordinate_file, stringsAsFactors = FALSE, fileEncoding = "UTF-8")
mean_monthly_folder <- "C:/Users/raza/project/mean_monthly_data"
file_list <- list.files(mean_monthly_folder, pattern = "\\.csv$", full.names = TRUE)
for (file in file_list) {
well_data <- read.csv(file, stringsAsFactors = FALSE)
merged_data <- merge(well_data, coordinates_data, by = "ID", all.x = TRUE)
write.csv(merged_data, paste0("merged_", basename(file)), row.names = FALSE)
}
Calculate mean monthly GWL
monthly_files <- list.files(destination_folder, pattern = "\\.csv$", full.names = TRUE)
monthly_mean_list <- list()
for (file in monthly_files) {
data <- read.csv(file)
data$Date <- as.Date(data$Date)
data$Month <- format(data$Date, "%Y-%m")
monthly_mean <- data %>%
group_by(ID, Month) %>%
summarize(Mean_GWL = mean(GWL, na.rm = TRUE), .groups = "drop")
monthly_mean_list[[basename(file)]] <- monthly_mean
write.csv(monthly_mean, gsub(".csv", "_monthly_mean.csv", file), row.names = FALSE)
}
Clean stations with missing months and transpose for PCA
clean_folder <- "C:/Users/raza/project/mean_monthly_data"
file_list <- list.files(clean_folder, pattern = "*.csv", full.names = TRUE)
for (file in file_list) {
data <- read.csv(file)
data$Date <- as.yearmon(data$Month, "%Y-%m")
ids_to_remove <- data %>%
group_by(ID) %>%
summarize(Num_Unique_Months = n_distinct(Date)) %>%
filter(Num_Unique_Months < 12) %>%
pull(ID)
data_filtered <- data %>% filter(!ID %in% ids_to_remove)
clean_filename <- paste0("clean_", gsub("[^0-9]", "", basename(file)), ".csv")
write.csv(data_filtered, clean_filename, row.names = FALSE)
data_transposed <- pivot_wider(data_filtered, names_from = ID, values_from = Mean_GWL)
data_transposed <- data_transposed %>% select(where(~ !any(is.na(.))))
transposed_filename <- paste0("transposed_", gsub("[^0-9]", "", basename(file)), ".csv")
write.csv(data_transposed, transposed_filename, row.names = TRUE)
}
PCA Analysis
data <- read.csv("transposed_2001-05.csv")
data_num <- data[, -1] # remove first column if necessary
data_num <- sapply(data_num, as.numeric)
data_num[is.na(data_num)] <- mean(data_num, na.rm = TRUE)
z_scaled_data <- scale(data_num)
pca_result <- prcomp(z_scaled_data)
summary(pca_result)
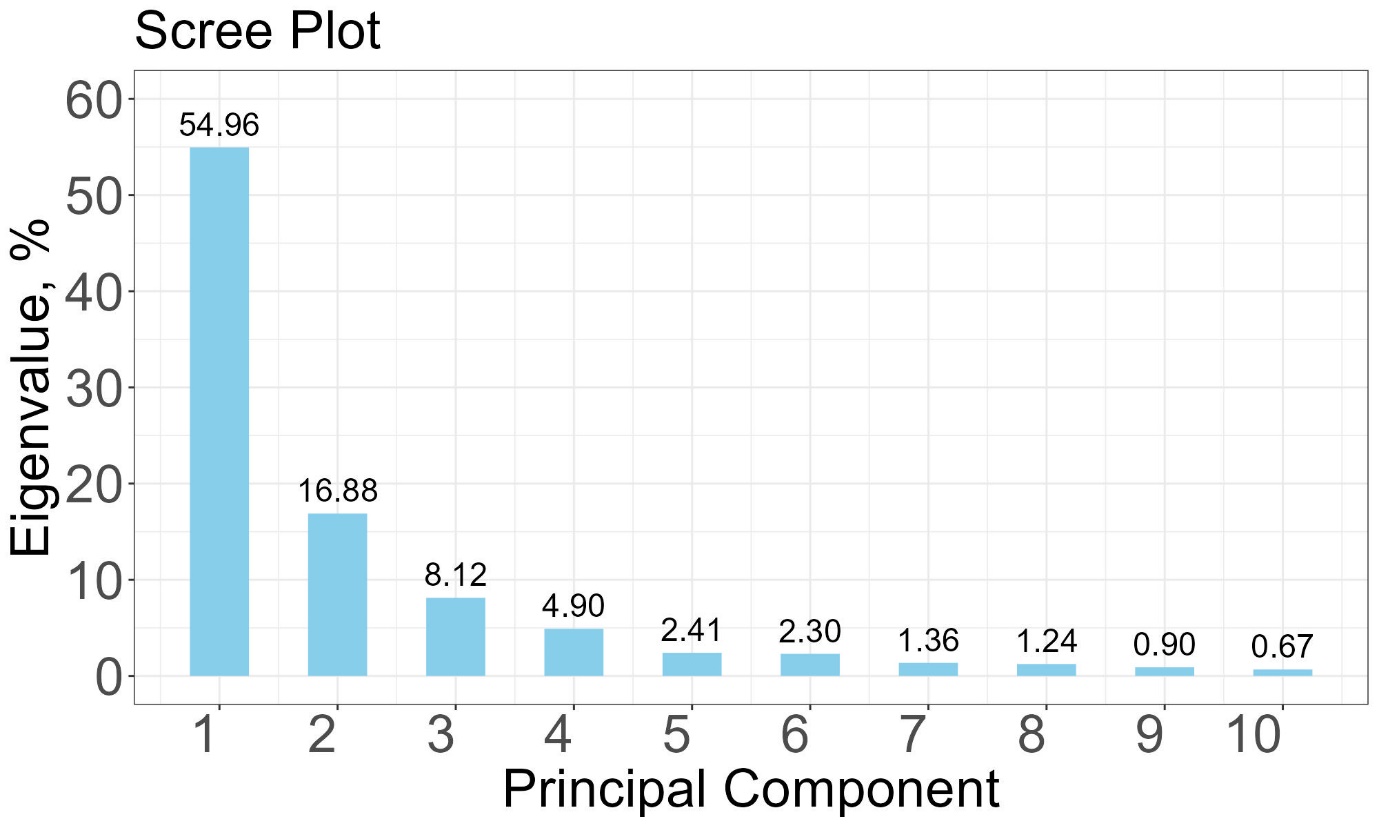
# Scree plot
eigenvalues <- (pca_result$sdev)^2 / sum((pca_result$sdev)^2) * 100
eig_df <- data.frame(Dimension = 1:length(eigenvalues), Eigenvalue = eigenvalues)
eig_df_sub <- head(eig_df, 10)
p <- ggplot(eig_df_sub, aes(x = Dimension, y = Eigenvalue)) +
geom_bar(stat = "identity", fill = "skyblue", width = 0.5) +
geom_text(aes(label = round(Eigenvalue, 2)), vjust = -0.5, size = 6) +
labs(title = "Scree Plot", x = "Principal Component", y = "Variance (%)") +
theme_bw() +
theme(axis.text = element_text(size = 28), axis.title = element_text(size = 28), plot.title = element_text(size = 28))
ggsave("scree_plot_1.jpg", plot = p, width = 10, height = 6, units = "in", dpi = 200)
# Save PCA results
write.csv(pca_result$x, "scores_2001-05.csv")
write.csv(pca_result$rotation, "loadings_2001-05.csv")
Reference Hydrograph & Residuals
pc_scores <- as.data.frame(pca_result$x)
regression_data <- cbind(data_num, pc_scores)
reference_hydrographs <- data.frame()
residuals_data <- data
for (well_col in 1:ncol(data_num)) {
lm_result <- lm(data_num[, well_col] ~ PC1 + PC2 + PC3 + PC4, data = regression_data)
reference <- predict(lm_result)
reference_hydrographs <- rbind(reference_hydrographs, data.frame(
Well = colnames(data_num)[well_col],
Reference_Hydrograph = reference
))
residuals_data[, well_col] <- data_num[, well_col] - reference
}
write.csv(reference_hydrographs, "reference_hydrographs.csv")
write.csv(residuals_data, "residuals_data.csv")
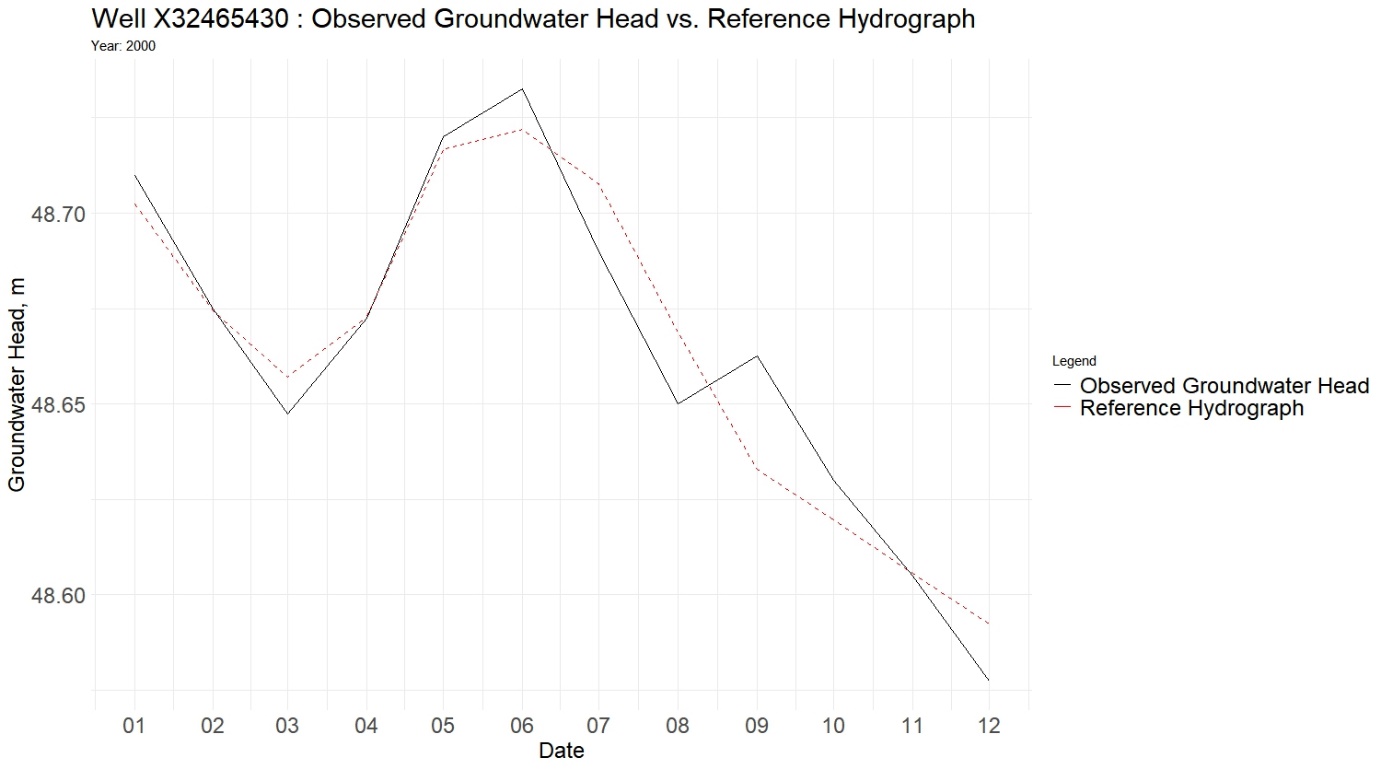
Plot Observed vs Reference Hydrograph
well_name <- "X32465430"
plot_data <- data.frame(
Date = as.Date(data$Month),
Groundwater_Head = data[[well_name]],
Reference_Hydrograph = reference_hydrographs$Reference_Hydrograph[reference_hydrographs$Well == well_name]
)
ggplot(plot_data, aes(x = Date)) +
geom_line(aes(y = Groundwater_Head, color = "Observed"), size = 1) +
geom_line(aes(y = Reference_Hydrograph, color = "Reference"), size = 1, linetype = "dashed") +
labs(title = paste("Well", well_name), x = "Year", y = "Groundwater Head") +
scale_color_manual(values = c("Observed" = "black", "Reference" = "red")) +
theme_bw()
Calculate RMSE
well_rmses <- data.frame(Well = colnames(data_num), RMSE = NA)
for (i in seq_along(well_rmses$Well)) {
obs <- data_num[, i]
ref <- reference_hydrographs$Reference_Hydrograph[reference_hydrographs$Well == well_rmses$Well[i]]
well_rmses$RMSE[i] <- sqrt(mean((obs - ref)^2))
}
ranked_wells <- well_rmses[order(well_rmses$RMSE), ]
write.csv(well_rmses, "RMSE_2001-05.csv")
plots with RMSE > 0.25m
setwd("C:/Users/raza/project")
data <- read.csv("total.csv")
plot <- ggplot(data_melt, aes(x = factor(year), y = value, color = variable, group = variable)) +
geom_line(size = 1.2) + # Line plot with increased line width
labs(title = "",
x = "Year",
y = "Number of wells") +
theme_bw() +
theme(legend.position = "bottom",
axis.text.x = element_text(angle = 0, hjust = 1, size = 16, face = "bold"),
axis.text.y = element_text(size = 16, face = "bold"),
axis.title = element_text(size = 18),
legend.title = element_blank(),
legend.text = element_text(size = 14, face = "bold"),
plot.title = element_text(size = 14, face = "bold")) +
scale_color_manual(values = c("Total.Wells" = "#E41A1C", "Complete.Data" = "#377EB8",
"RMSE...0.25m" = "darkgreen", "RMSE....0.25m" = "darkgoldenrod",
"Missing.Data" = "orange"),
labels = c("Total Wells",
"Wells with complete data",
"RMSE <= 0.25m",
"Wells with missing data",
"RMSE > 0.25m")) +
scale_x_discrete(breaks = seq(2001, 2005, by = 1)) +
geom_vline(xintercept = c("2001", "2005"),
linetype = "dashed", color = "black", size = 0.45) +
guides(color = guide_legend(nrow = 2))
plot
# Save the plot as a PNG file
ggsave("total_v2.png", plot = plot, width = 10, height = 7, dpi = 100)