Session 1
Technical setup Data, scripts/notebooks, environment and folder structure needed for the practical sessions are available in the folder Workshop-MSRS4A_elearning.zip, accessed through the link https://github.com/zalf-rpm/KIKompAg/tree/main/Lesson1
Please follow instructions provided below and attached in the readme.html file.
Planet-Scope_Sentinel-2 Data Fusion with Random Forest
In this tutorial we show how to train the RF model and generate synthetic Sentinel-2 (S2) images, based on Planet-Scope (PS) data from 2021 and 2022. The validation is exemplified in the maps comparing original S2, synthetic S2 and resampled PS images, as well as absolute error synthetic vs original S2 and SSIM index are compared.
Input:
- Sentinel-2 NDVI images
- PlanetScope NDVI images
- IACS crop type vectors
Outputs:
- Output folder named <{Dataset_Fused}>
- Synthetic S2 images named Synth-rf_ndvidoy<{year}>_<{planetdoy:03d}>.tif
Steps:
- Global settings
- Reading the S2 and PS NDVI dataset and maize mask for both years
- Definition of matching image pairs and application
- Creating feature matrix and processing in loop
- Training RF model
- Visualisation of RF-based and resampling-based synthetic S2 outputs
- Loading and merging real S2 and PS images
- Applying the RF-model and saving synthetic S2 images
Main script with basis functions is PS_S2_fusion.py
1. Global settings and Python framework
# Import the Python frameworks
from PS_S2_fusion import (
rasterio,
NDVIDataLoader,
NDVIProcessor,
random,
Dict,
NDVIImage,
np,
# LinearRegression,
RandomForestRegressor,
NDVIVisualizer,
)
from sklearn.model_selection import train_test_split
from sklearn.metrics import mean_squared_error
import pprint
from pathlib import Path
import copy
from skimage.metrics import structural_similarity as ssim
import numpy as np# Global flag: Set to True to include before/after Sentinel-2 images as features, False to exclude
INCLUDE_BEFORE_AFTER = False
random_state=42
np.set_printoptions(formatter={'float': '{: 0.3f}'.format})
# Input and output dir settings
input_dir = Path("./Dataset_original/13/")# Saving function
def save_tif(data, output_file, crs, transform):
"""Saves raster data to a GeoTIFF file.
Args:
data (array-like): The 2D or 3D raster data to save.
output_file (str): The path and filename of the output GeoTIFF file.
crs (str or rasterio.crs.CRS): The Coordinate Reference System.
Can be an EPSG code string (e.g., 'EPSG:4326')
or a rasterio.crs.CRS object.
transform (affine.Affine): The affine transformation for the raster.
Returns:
None
"""
# Determine number of bands
if data.ndim == 2:
count = 1
elif data.ndim == 3:
count = data.shape[0] # Assume bands are in the first dimension
else:
raise ValueError("Data must be 2D or 3D.")
profile = {
"driver": "GTiff",
"height": data.shape[1] if data.ndim == 3 else data.shape[0],
"width": data.shape[2] if data.ndim == 3 else data.shape[1],
"count": count,
"dtype": data.dtype.name, # Use data type of input data
"crs": crs,
"transform": transform,
}
# Ensure directory exists before saving
output_file.parent.mkdir(parents=True, exist_ok=True)
with rasterio.open(output_file, "w", **profile) as dst:
if count == 1:
dst.write(data, 1)
else:
for band_index in range(1, count + 1):
dst.write(data[band_index - 1], band_index)2. Reading the S2 and PS NDVI dataset and maize mask for both years
# Reading NDVI data
data_loader = NDVIDataLoader()
processor = NDVIProcessor()
shapefile_2021 = input_dir / "2021/shp/2021_fid13_maize_subset.shp"
shapefile_2022 = input_dir / "2022/shp/2022_fid13_maize_subset.shp"
sentinel_2_images_2021 = data_loader.load_ndvi_images(
input_dir / "2021/ndvi/S2_maize/", "S2*.tif"
)
planet_scope_images_2021 = data_loader.load_ndvi_images(
input_dir / "2021/ndvi/PS_maize/", "PS*.tif"
)
sentinel_2_images_2022 = data_loader.load_ndvi_images(
input_dir / "2022/ndvi/S2_maize/", "S2*.tif"
)
planet_scope_images_2022 = data_loader.load_ndvi_images(
input_dir / "2022/ndvi/PS_maize/", "PS*.tif"
)
print('DONE: Reading Sentinel-2 and PlanetScope images')3. Definition of matching image pairs and application
def find_matching_image_pairs(sentinel_images, planet_images, year):
"""
Finds matching image pairs from Sentinel-2 and PlanetScope dictionaries
based on DOY with a descriptive output.
Args:
sentinel_images: Dictionary with DOY as key and Sentinel-2 NDVI data as value.
planet_images: Dictionary with DOY as key and PlanetScope NDVI data as value.
Returns:
A list of dictionaries, where each dictionary represents a match:
{'planet_doy': ...,
'sentinel_matching_doy': ...,
'sentinel_before_doy': ...,
'sentinel_after_doy': ...}
"""
matching_pairs = []
for planet_doy in planet_images:
for doy_offset in range(-1, 2): # Check DOY ± 3 days
matching_sentinel_doy = planet_doy + doy_offset
if matching_sentinel_doy in sentinel_images:
# Find nearest before/after Sentinel images
sentinel_before_doy = max(
[doy for doy in sentinel_images if doy < matching_sentinel_doy]
)
sentinel_after_doy = min(
[doy for doy in sentinel_images if doy > matching_sentinel_doy]
)
if INCLUDE_BEFORE_AFTER:
matching_pairs.append(
{
"year": year,
"planet_doy": planet_doy,
"sentinel_matching_doy": matching_sentinel_doy,
"sentinel_before_doy": sentinel_before_doy,
"sentinel_after_doy": sentinel_after_doy,
}
)
else:
matching_pairs.append(
{
"year": year,
"planet_doy": planet_doy,
"sentinel_matching_doy": matching_sentinel_doy,
}
)
break # Move on to the next PlanetScope image after a match
return matching_pairs# Applying matching function to our dataset:
matching_pairs_2022 = find_matching_image_pairs(
sentinel_2_images_2022, planet_scope_images_2022, 2022
)
matching_pairs_2021 = find_matching_image_pairs(
sentinel_2_images_2021, planet_scope_images_2021, 2021
)
print('DONE: Matching pairs found')
pprint.pp(matching_pairs_2021)
pprint.pp(matching_pairs_2022)DONE: Matching pairs found
[{'year': 2021, 'planet_doy': 111, 'sentinel_matching_doy': 111},
{'year': 2021, 'planet_doy': 117, 'sentinel_matching_doy': 118},
{'year': 2021, 'planet_doy': 150, 'sentinel_matching_doy': 151},
{'year': 2021, 'planet_doy': 151, 'sentinel_matching_doy': 151},
{'year': 2021, 'planet_doy': 153, 'sentinel_matching_doy': 153},
{'year': 2021, 'planet_doy': 167, 'sentinel_matching_doy': 168},
{'year': 2021, 'planet_doy': 168, 'sentinel_matching_doy': 168},
{'year': 2021, 'planet_doy': 169, 'sentinel_matching_doy': 168},
{'year': 2021, 'planet_doy': 225, 'sentinel_matching_doy': 226},
{'year': 2021, 'planet_doy': 226, 'sentinel_matching_doy': 226},
{'year': 2021, 'planet_doy': 227, 'sentinel_matching_doy': 226},
{'year': 2021, 'planet_doy': 251, 'sentinel_matching_doy': 251},
{'year': 2021, 'planet_doy': 280, 'sentinel_matching_doy': 281},
{'year': 2021, 'planet_doy': 282, 'sentinel_matching_doy': 281}]
[{'year': 2022, 'planet_doy': 113, 'sentinel_matching_doy': 113},
{'year': 2022, 'planet_doy': 114, 'sentinel_matching_doy': 113},
{'year': 2022, 'planet_doy': 122, 'sentinel_matching_doy': 123},
{'year': 2022, 'planet_doy': 123, 'sentinel_matching_doy': 123},
{'year': 2022, 'planet_doy': 138, 'sentinel_matching_doy': 138},
{'year': 2022, 'planet_doy': 166, 'sentinel_matching_doy': 166},
{'year': 2022, 'planet_doy': 193, 'sentinel_matching_doy': 193},
{'year': 2022, 'planet_doy': 221, 'sentinel_matching_doy': 221},
{'year': 2022, 'planet_doy': 222, 'sentinel_matching_doy': 221},
{'year': 2022, 'planet_doy': 248, 'sentinel_matching_doy': 248},
{'year': 2022, 'planet_doy': 273, 'sentinel_matching_doy': 273}]# Combining matching pairs from both years for training the RF model and validation:
matching_pairs = matching_pairs_2021 + matching_pairs_2022
validation_pairs = random.sample(matching_pairs, 5)
training_pairs = [
element for element in matching_pairs if element not in validation_pairs
]
print('DONE: Random selection of pairs for training, testing and validation')
pprint.pp(validation_pairs)
len(training_pairs)DONE: Random selection of pairs for training, testing and validation
[{'year': 2021, 'planet_doy': 153, 'sentinel_matching_doy': 153},
{'year': 2022, 'planet_doy': 113, 'sentinel_matching_doy': 113},
{'year': 2021, 'planet_doy': 151, 'sentinel_matching_doy': 151},
{'year': 2021, 'planet_doy': 111, 'sentinel_matching_doy': 111},
{'year': 2022, 'planet_doy': 166, 'sentinel_matching_doy': 166}]204. Creating feature matrix and processing in loop
# Loading the tiff (feature = input variables)
def create_feature_target_pair(
match,
sentinel_images: Dict[int, NDVIImage],
planet_images: Dict[int, NDVIImage],
shapefile_path: Path,
):
"""
Creates a single feature array (X) and target array (y) pair
for one set of matching images.
Args:
match: A single dictionary from the matching_pairs list.
sentinel_images: Dictionary with DOY as key and Sentinel-2 NDVIImage.
planet_images: Dictionary with DOY as key and PlanetScope NDVIImage.
Returns:
A tuple: (features (X), targets (y))
- features (X): 2D NumPy array of features, one row per valid pixel.
- targets (y): 1D NumPy array of corresponding Sentinel-2 NDVI.
"""
processor = NDVIProcessor()
year = match["year"]
planet_doy = match["planet_doy"]
sentinel_matching_doy = match["sentinel_matching_doy"]
if INCLUDE_BEFORE_AFTER:
sentinel_before_doy = match["sentinel_before_doy"]
sentinel_after_doy = match["sentinel_after_doy"]
# Resample PlanetScope
planet_image = processor.resample_planet_to_sentinel(
sentinel_images[sentinel_matching_doy], planet_images[planet_doy]
)
# Extract pixel data
sentinel_matching_image = sentinel_images[sentinel_matching_doy]
if INCLUDE_BEFORE_AFTER:
sentinel_before_image = sentinel_images[sentinel_before_doy]
sentinel_after_image = sentinel_images[sentinel_after_doy]
planet_image.ndvi = processor.mask_array_with_shapefile(
planet_image.ndvi, planet_image.transform, shapefile_path
)
sentinel_matching_image.ndvi = processor.mask_array_with_shapefile(
sentinel_matching_image.ndvi, sentinel_matching_image.transform, shapefile_path
)
if INCLUDE_BEFORE_AFTER:
sentinel_before_image.ndvi = processor.mask_array_with_shapefile(
sentinel_before_image.ndvi, sentinel_before_image.transform, shapefile_path
)
sentinel_after_image.ndvi = processor.mask_array_with_shapefile(
sentinel_after_image.ndvi, sentinel_after_image.transform, shapefile_path
)
# Processing with invalid_mask as before
invalid_mask_matching = processor.get_invalid_mask(
sentinel_ndvi_data=sentinel_matching_image.ndvi,
planet_ndvi_data=planet_image.ndvi,
)
invalid_mask = invalid_mask_matching # for +/-1 day offset
if INCLUDE_BEFORE_AFTER:
invalid_mask_before_after = processor.get_invalid_mask(
sentinel_before_image.ndvi, sentinel_after_image.ndvi
)
invalid_mask = np.logical_or(invalid_mask_matching, invalid_mask_before_after)
sentinel_matching_data = processor.get_preprocessed_ndvi_data(
sentinel_matching_image, invalid_mask
).flatten()
if INCLUDE_BEFORE_AFTER:
sentinel_before_data = processor.get_preprocessed_ndvi_data(
sentinel_before_image, invalid_mask
).flatten()
sentinel_after_data = processor.get_preprocessed_ndvi_data(
sentinel_after_image, invalid_mask
).flatten()
planet_data = processor.get_preprocessed_ndvi_data(
planet_image, invalid_mask
).flatten()
num_valid_pixels = len(sentinel_matching_data)
# Create feature matrix (one row per valid pixel)
if INCLUDE_BEFORE_AFTER:
features = np.column_stack(
(
planet_data,
sentinel_before_data,
sentinel_after_data,
np.full(num_valid_pixels, planet_doy),
np.full(num_valid_pixels, sentinel_before_doy),
np.full(num_valid_pixels, sentinel_after_doy),
np.full(num_valid_pixels, year),
)
)
else:
features = np.column_stack(
(
planet_data,
np.full(num_valid_pixels, planet_doy),
np.full(num_valid_pixels, year),
)
)
targets = sentinel_matching_data
return features, targets, invalid_mask# Processing data for each match in a loop
all_features = []
all_targets = []
all_masks = []
for match in training_pairs:
if match["year"] == 2022:
X, y, invalid_mask = create_feature_target_pair(
match,
sentinel_2_images_2022,
planet_scope_images_2022,
shapefile_2022,
)
else:
X, y, invalid_mask = create_feature_target_pair(
match,
sentinel_2_images_2021,
planet_scope_images_2021,
shapefile_2021,
)
all_features.append(X)
all_targets.append(y)
all_masks.append(invalid_mask)
# Concatenate the results if needed
X = np.concatenate(all_features, axis=0)
y = np.concatenate(all_targets)
print(X.shape, y.shape)
print(X)
print(y)(774349, 3) (774349,)
[[ 0.325 117.000 2021.000]
[ 0.280 117.000 2021.000]
[ 0.310 117.000 2021.000]
...
[ 0.445 273.000 2022.000]
[ 0.627 273.000 2022.000]
[ 0.518 273.000 2022.000]]
[ 0.346 0.255 0.221 ... 0.146 0.306 0.314]5. Training RF model
# Using RF model with following settings: n_jobs= performance (parallelisation); n_estimators= The number of trees in the forest;
# max_depth= The maximum depth of the tree; random_state = 42 (= reproducible); 20% Testing, 80% Training
# Description: https://scikit-learn.org/stable/modules/generated/sklearn.ensemble.RandomForestRegressor.html
rf_model = RandomForestRegressor(
n_estimators=40, max_depth=15, random_state=42, n_jobs=8
)
X_train, X_test, y_train, y_test = train_test_split(
X, y, test_size=0.2, random_state=42
)
rf_model.fit(X_train, y_train)
predictions = rf_model.predict(X_test)
mse = mean_squared_error(y_test, predictions)
print(f'Mean squared error: {mse}')Mean squared error: 0.0007536957916862879
6. Visualisation of RF-based and resampling-based synthetic S2 outputs
# Printing validation pairs
pprint.pp(validation_pairs)[{'year': 2021, 'planet_doy': 153, 'sentinel_matching_doy': 153},
{'year': 2022, 'planet_doy': 113, 'sentinel_matching_doy': 113},
{'year': 2021, 'planet_doy': 151, 'sentinel_matching_doy': 151},
{'year': 2021, 'planet_doy': 111, 'sentinel_matching_doy': 111},
{'year': 2022, 'planet_doy': 166, 'sentinel_matching_doy': 166}]## Visualisation setup
# Precompute features, targets, masks, and predictions for all validation pairs
all_features = []
all_targets = []
all_masks = []
predictions_list = []
for match in validation_pairs:
year = match["year"]
if year == 2021:
X, y, invalid_mask = create_feature_target_pair(
match, sentinel_2_images_2021, planet_scope_images_2021, shapefile_2021
)
else:
X, y, invalid_mask = create_feature_target_pair(
match, sentinel_2_images_2022, planet_scope_images_2022, shapefile_2022
)
all_features.append(X)
all_targets.append(y)
all_masks.append(invalid_mask)
# Precompute prediction
pred = rf_model.predict(X)
predictions_list.append(pred)
# Select one match for 2021 and one for 2022 within DOY 160–260
data_to_visualize = []
for year in (2021, 2022):
found = next(
(m for m in validation_pairs
if m["year"] == year and 150 <= m["planet_doy"] <= 260),
None
)
if found:
data_to_visualize.append(found)
# Visualization loop over the two selected matches
mse_dict_synth = {}
mse_dict_resampled = {}
mean_ndvi_dict_synth = {}
mean_ndvi_dict_resampled = {}
mean_ndvi_dict_real = {}
visualizer = NDVIVisualizer()
for match in data_to_visualize:
# find corresponding index i safely
i = next(
idx for idx, m in enumerate(validation_pairs)
if m["year"] == match["year"]
and m["planet_doy"] == match["planet_doy"]
and m["sentinel_matching_doy"] == match["sentinel_matching_doy"]
)
# pick data and images
year = match["year"]
if year == 2021:
s2_images = sentinel_2_images_2021
ps_images = planet_scope_images_2021
shapefile = shapefile_2021
else:
s2_images = sentinel_2_images_2022
ps_images = planet_scope_images_2022
shapefile = shapefile_2022
planet_doy = match["planet_doy"]
sentinel_matching_doy = match["sentinel_matching_doy"]
preds = predictions_list[i]
# Create synthetic NDVI image
synth_image = copy.deepcopy(s2_images[sentinel_matching_doy])
synth_image.ndvi[~all_masks[i]] = preds
# Resample PlanetScope and mask
resampled = processor.resample_planet_to_sentinel(
s2_images[sentinel_matching_doy],
ps_images[planet_doy],
)
resampled.ndvi = processor.mask_array_with_shapefile(
resampled.ndvi, resampled.transform, shapefile
)
# Visualize
visualizer.visualize_ndvi_images(
s2_images[sentinel_matching_doy].ndvi,
synth_image.ndvi,
resampled.ndvi,
planet_doy,
year,
)
# Compute metrics
mse_synth = mean_squared_error(all_targets[i], preds)
mse_resampled = mean_squared_error(
all_targets[i],
processor.get_preprocessed_ndvi_data(resampled, all_masks[i]),
)
ssim_score = ssim(
np.nan_to_num((s2_images[sentinel_matching_doy].ndvi - np.nanmin(
s2_images[sentinel_matching_doy].ndvi)) /
(np.nanmax(s2_images[sentinel_matching_doy].ndvi) -
np.nanmin(s2_images[sentinel_matching_doy].ndvi))),
np.nan_to_num((synth_image.ndvi - np.nanmin(synth_image.ndvi)) /
(np.nanmax(synth_image.ndvi) -
np.nanmin(synth_image.ndvi))),
data_range=1.0,
)
print(f"=== Year {year}, DOY {planet_doy} ===")
print(f"MSE synthetic: {mse_synth:.5f}")
print(f"MSE resampled: {mse_resampled:.5f}")
print(f"SSIM (synthetic vs real): {ssim_score:.5f}")
# Store summary stats
mse_dict_synth[planet_doy] = mse_synth
mse_dict_resampled[planet_doy] = mse_resampled
mean_ndvi_dict_synth[planet_doy] = np.mean(preds)
mean_ndvi_dict_resampled[planet_doy] = np.nanmean(resampled.ndvi)
mean_ndvi_dict_real[planet_doy] = np.nanmean(
s2_images[sentinel_matching_doy].ndvi
)
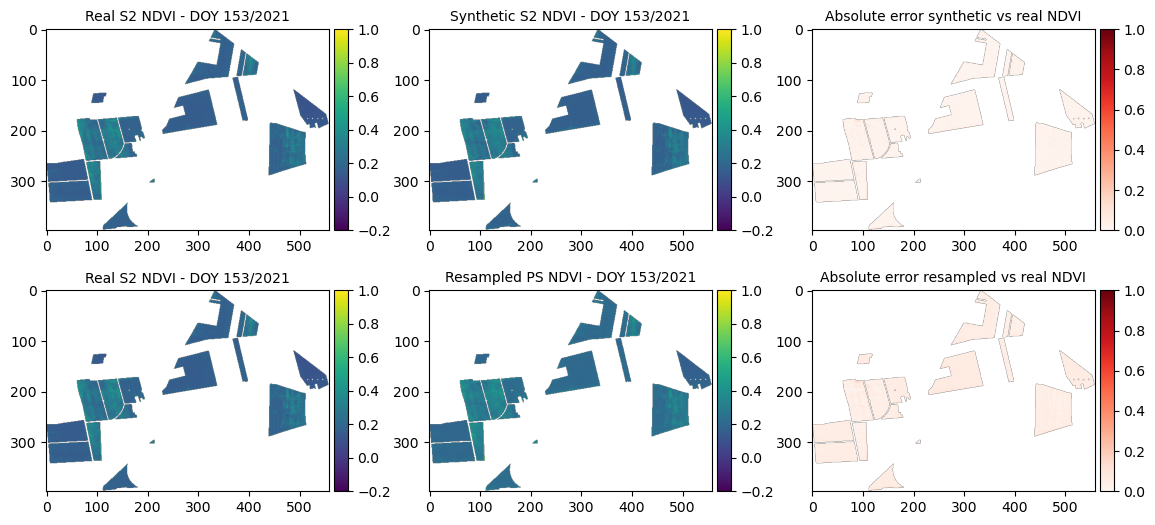
=== Year 2021, DOY 153 ===
MSE synthetic: 0.00045
MSE resampled: 0.00255
SSIM (synthetic vs real): 0.96706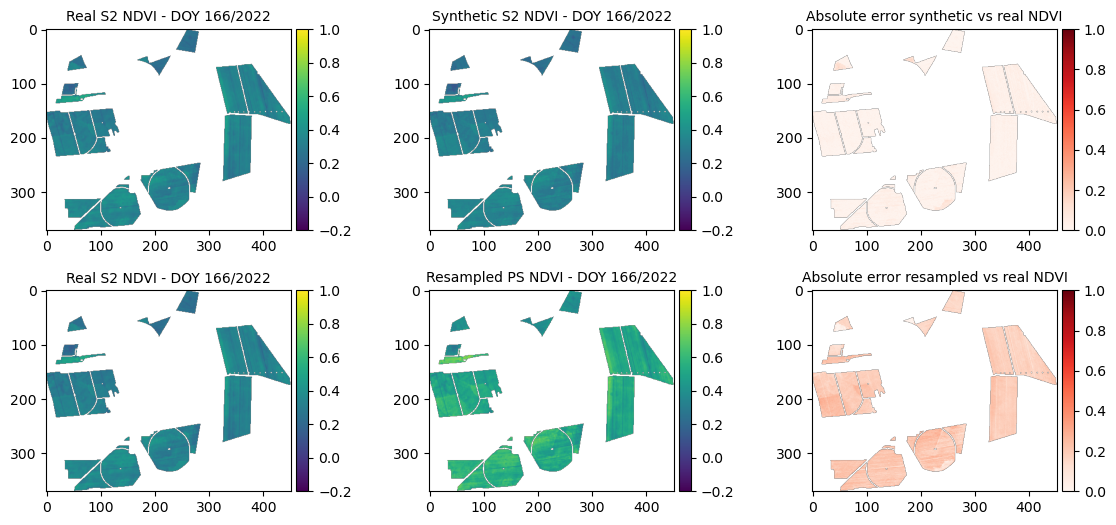
=== Year 2021, DOY 153 ===
MSE synthetic: 0.00045
MSE resampled: 0.00255
SSIM (synthetic vs real): 0.96706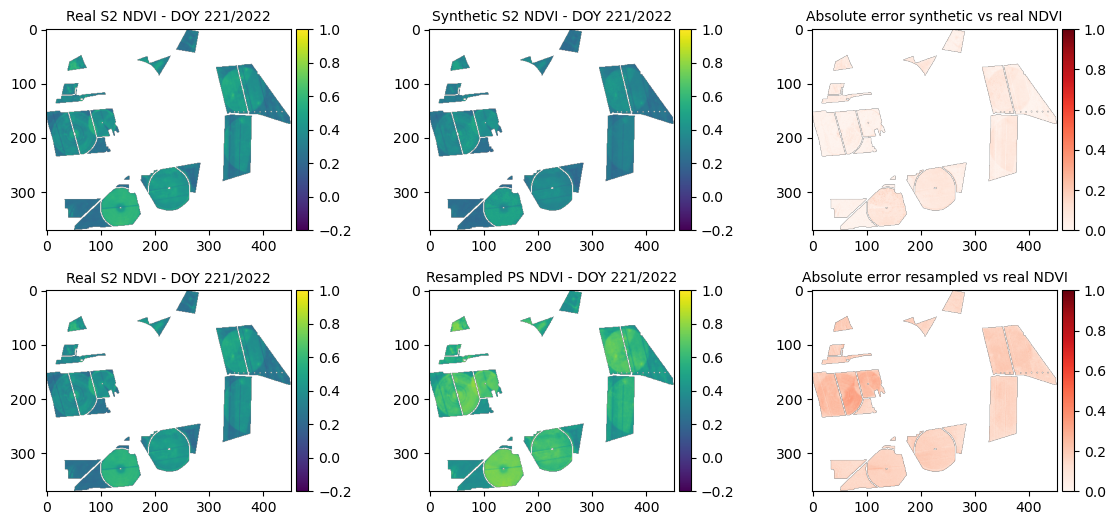
print(f"Total Error Synthetic: {sum(mse_dict_synth.values())}")
print(f"Total Error Resampled: {sum(mse_dict_resampled.values())}")7. Loading and merging real S2 and PS images
# Generate real matching pairs
def find_closest_sentinel_images(sentinel_images, planet_images, year):
"""
Finds matching image pairs from Sentinel-2 and PlanetScope dictionaries
based on DOY with a descriptive output.
Args:
sentinel_images: Dictionary with DOY as key and Sentinel-2 NDVI data as value.
planet_images: Dictionary with DOY as key and PlanetScope NDVI data as value.
Returns:
A list of dictionaries, where each dictionary represents a match:
{'planet_doy': ...,
'sentinel_before_doy': ...,
'sentinel_after_doy': ...}
"""
set_of_input_images = []
for planet_doy in planet_images.keys():
## Find nearest before/after Sentinel images
sentinel_before_doy = max([doy for doy in sentinel_images if doy < planet_doy])
sentinel_after_doy = min([doy for doy in sentinel_images if doy > planet_doy])
set_of_input_images.append(
{
"year": year,
"planet_doy": planet_doy,
"sentinel_before_doy": sentinel_before_doy,
"sentinel_after_doy": sentinel_after_doy,
}
)
return set_of_input_images
# Get all input images
set_of_input_images_2021 = find_closest_sentinel_images(
sentinel_2_images_2021, planet_scope_images_2021, 2021
)
set_of_input_images_2022 = find_closest_sentinel_images(
sentinel_2_images_2022, planet_scope_images_2022, 2022
)
# Generate all input features
def create_input_data(
match,
sentinel_images: Dict[int, NDVIImage],
planet_images: Dict[int, NDVIImage],
shapefile_path: Path,
):
"""
Creates a single feature array (X) and target array (y) pair
for one set of matching images.
Args:
match: A single dictionary from the matching_pairs list.
sentinel_images: Dictionary with DOY as key and Sentinel-2 NDVIImage.
planet_images: Dictionary with DOY as key and PlanetScope NDVIImage.
Returns:
A tuple: (features (X), targets (y))
- features (X): 2D NumPy array of features, one row per valid pixel.
- targets (y): 1D NumPy array of corresponding Sentinel-2 NDVI.
"""
processor = NDVIProcessor()
year = match["year"]
planet_doy = match["planet_doy"]
sentinel_before_doy = match["sentinel_before_doy"]
sentinel_after_doy = match["sentinel_after_doy"]
# Resample PlanetScope
planet_image = processor.resample_planet_to_sentinel(
sentinel_images[sentinel_before_doy], planet_images[planet_doy]
)
# Extract pixel data
sentinel_before_image = sentinel_images[sentinel_before_doy]
sentinel_after_image = sentinel_images[sentinel_after_doy]
# Include maize
planet_image.ndvi = processor.mask_array_with_shapefile(
planet_image.ndvi, planet_image.transform, shapefile_path
)
sentinel_before_image.ndvi = processor.mask_array_with_shapefile(
sentinel_before_image.ndvi, sentinel_before_image.transform, shapefile_path
)
sentinel_after_image.ndvi = processor.mask_array_with_shapefile(
sentinel_after_image.ndvi, sentinel_after_image.transform, shapefile_path
)
invalid_mask_matching = processor.get_invalid_mask(
sentinel_ndvi_data=sentinel_before_image.ndvi,
planet_ndvi_data=planet_image.ndvi,
)
invalid_mask_before_after = processor.get_invalid_mask(
sentinel_before_image.ndvi, sentinel_after_image.ndvi
)
invalid_mask = np.logical_or(invalid_mask_matching, invalid_mask_before_after)
invalid_mask = invalid_mask_matching
if INCLUDE_BEFORE_AFTER:
sentinel_before_data = processor.get_preprocessed_ndvi_data(
sentinel_before_image, invalid_mask
).flatten()
sentinel_after_data = processor.get_preprocessed_ndvi_data(
sentinel_after_image, invalid_mask
).flatten()
planet_data = processor.get_preprocessed_ndvi_data(
planet_image, invalid_mask
).flatten()
num_valid_pixels = (
len(sentinel_before_data) if INCLUDE_BEFORE_AFTER else len(planet_data)
)
# Create feature matrix (one row per valid pixel)
if INCLUDE_BEFORE_AFTER:
features = np.column_stack(
(
planet_data,
sentinel_before_data,
sentinel_after_data,
np.full(num_valid_pixels, planet_doy),
np.full(num_valid_pixels, sentinel_before_doy),
np.full(num_valid_pixels, sentinel_after_doy),
np.full(num_valid_pixels, year),
)
)
else:
features = np.column_stack(
(
planet_data,
np.full(num_valid_pixels, planet_doy),
np.full(num_valid_pixels, year),
)
)
return features, invalid_mask# Combine and mask all input images
input_images = set_of_input_images_2021 + set_of_input_images_2022
# Process data for each match in a loop
all_inputs = []
all_masks = []
for match in input_images:
if match["year"] == 2022:
X, invalid_mask = create_input_data(
match,
sentinel_2_images_2022,
planet_scope_images_2022,
shapefile_2022,
)
else:
X, invalid_mask = create_input_data(
match,
sentinel_2_images_2021,
planet_scope_images_2021,
shapefile_2021,
)
all_inputs.append(X)
all_masks.append(invalid_mask)
print('DONE: Loading data and masking')DONE: Loading data and masking8. Applying the RF-model and saving synthetic S2 images
# Run RF-model and save all Synthetic NDVI images in sub-folder per year
base_output_dir = Path("./Dataset_Fused")
for i, match in enumerate(input_images):
year = match["year"]
if year == 2022:
sentinel_2_images = sentinel_2_images_2022
planet_scope_images = planet_scope_images_2022
else:
sentinel_2_images = sentinel_2_images_2021
planet_scope_images = planet_scope_images_2021
planet_doy = match["planet_doy"]
sentinel_before_doy = match["sentinel_before_doy"]
# Only synthesize if there is NOT a Sentinel-2 image for this DOY
if planet_doy not in sentinel_2_images:
# Make predictions
predictions = rf_model.predict(all_inputs[i])
# Use the correct reference image for output
if INCLUDE_BEFORE_AFTER:
synth_image = copy.deepcopy(sentinel_2_images[sentinel_before_doy])
else:
# If not using before/after, still need a reference image; use before_doy as default
synth_image = copy.deepcopy(sentinel_2_images[sentinel_before_doy])
synth_image.ndvi[~all_masks[i]] = predictions
# Create year-specific subfolder inside output_dir
year_output_dir = base_output_dir / str(year)
output_path = year_output_dir / f"Synth-rf_ndvi_doy_{year}_{planet_doy:03d}_.tif"
save_tif(
synth_image.ndvi,
output_path,
synth_image.crs,
synth_image.transform,
)
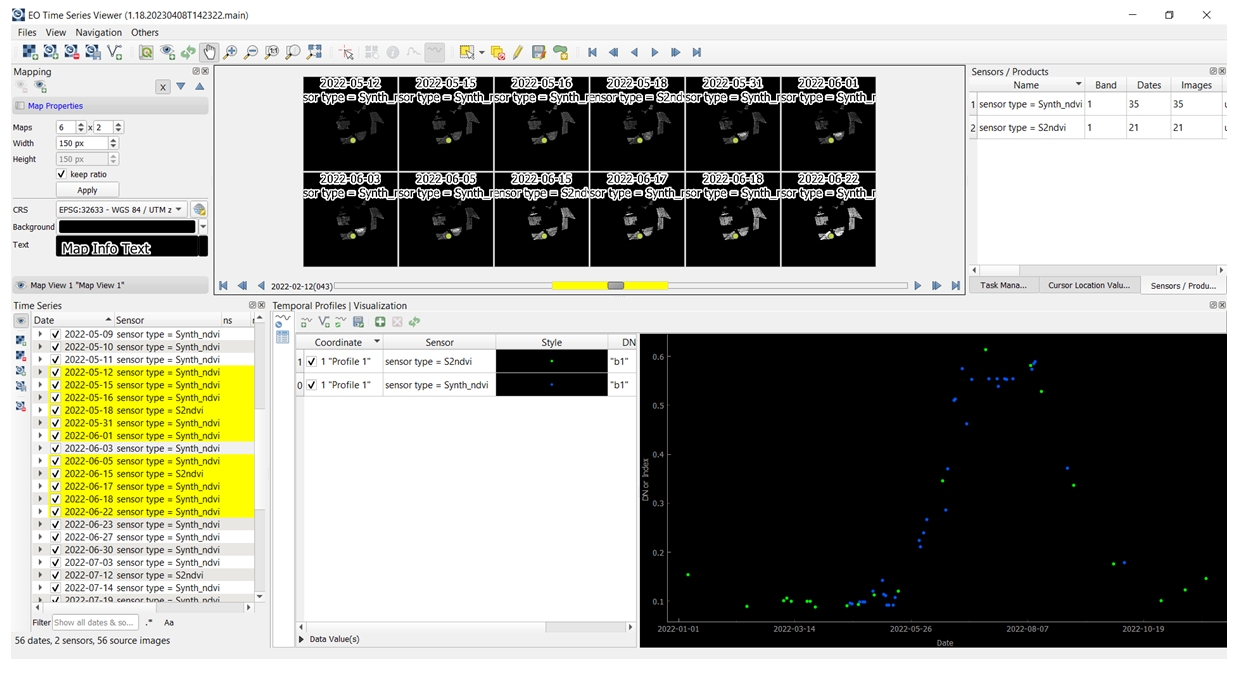
print(f'DONE: All synthetic data saved to {base_output_dir}')DONE: All synthetic data saved to Dataset_FusedOptional: After the RF model was trained and synthetic S2 data created, one can upload the original and synthetic S2 NDVI time series in the QGIS EO Time Series Viewer application. To do so, please download and install QGIS 3.28 or update with the EO Time Series Viewer plugin. For more info on how to proceed check Documentation: https://eo-time-series-viewer.readthedocs.io/en/latest.
For this task, please drag & drop the original S2 NDVI images (S2_ndvi_doyDOY.tif) and synthetic S2 NDVI images (Synth-rf_ndvi_doy_2022DOY.tif) available in the following folder:
..\Hands-on\Practical_part_1\extras\ Dataset_Fused_2022_timesteps\ to the Time series data panel in the EO Time Series Viewer. Then in the Mapping panel set the number, size and zoom of single maps subset. In the next step, using Select temporal profile from map button collect the temporal profile from selected pixel and add it to the Temporal profile panel. Finally, under Sensor and Style in the same panel, the predefined sensor (S2ndvi or Synth_ndvi) and the style (different colours eg. green and blue dots respectively) can be selected. The figure below shows an example how synthetic S2 NDVI data (blue dots) fill the gaps within the time series of the original S2 NDVI data (green dots).