Action
Our case study field is located Booßen, a city located in east Germany, close to Poland border. The data is kindly provided by the project Präzise Kalkung in Brandenburg, or ph-BB, funded by:
- Ministerium für Ländliche Entwicklung, Umwelt und Landwirtschaft des Landes Brandenburg,
- Investitionsbank des Landes Brandenburg,
- Europäische Union - Europäischer Landwirtschaftsfonds für die Entwicklung des ländlichen Raums.
More information about the project can be found in: https://www.atb-potsdam.de/en/research/research-projects/project/projekt/ph-bb
Step 1:
The first step is to select the places where we are going to collect the data in our field.
Inside the WF, we find the folder ./data/_data_to_select_in_QGIS/, which contains the files required for the selection of location, all are maps in EPSG: 32633, including:
area.shp– containing the shape of the field (vector),bsi_2020-09-20.tif– the BSI index of the bare soil in a day where the field had its bare soil exposed (raster),ndwi_2021-06-22.tif– the NDWI when the vegetation over the field was close to its peak (raster),DEM.tif– the digital elevation map (raster).
Based on these files, we select extremes regarding the intensity of the quantities in each file. For this tutorial, the selected points are shown in the following figure
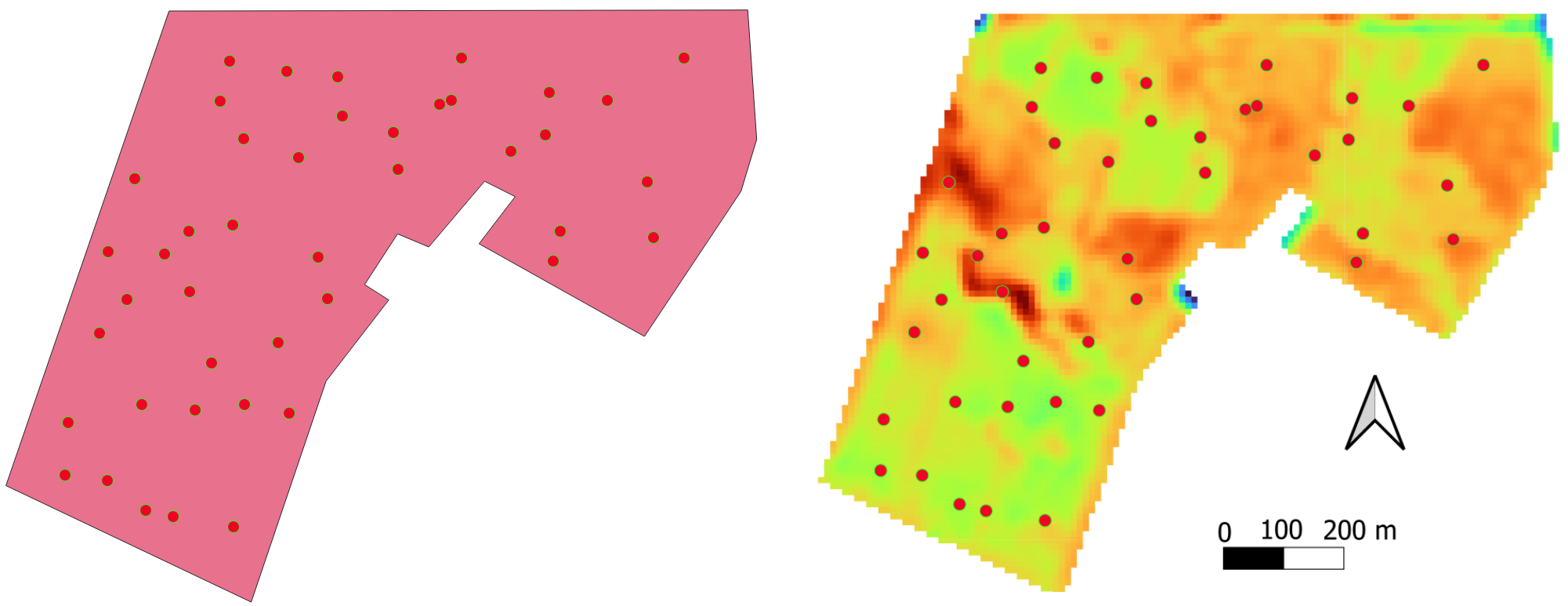
For each selected point, we will simulate that the farmer collects soil profile samples and crop yield at these specific points. The soil samples are submitted to laboratory analysis and the soil organic carbon, and the texture (sand, silt and clay contents) are known.
The coordinates of the selected points are then put into the file ./data/true_data/selected_coordinates.csv.
Step 2:
All the raw satellite images, already filtered by cloud percentage, are available in the folder ./data/all_imags_raw/. The images containing a date in the name, followed by “.tif” (such as in S2_2019-01-14.tif) are image data containing all the bands from Sentinel-2 S2L. The files ending in SCL.tif are the Scene Classification Layer (SCL). The codes for the SCL are available in:
https://custom-scripts.sentinel-hub.com/custom-scripts/sentinel-2/scene-classification/
At this stage, we are ready to run the first Python script, which aims to plot the temporal interpolation in a graph: ./py_codes/1.1_imag_reader.py. At the beginning of the scripts available in this course, all the configuration are setup.
In the configuration section, inside the script 1.1, we may select pixel coordinates (variables px_x and px_y) used to interpolate the temporal data (respective to each selected location). The variable lambda_whittaker is the lambda in the Whittaker function, responsible for increasing or reducing the smoothness. Both NDWI and BSI can be plotted by switching to True the respective variable ndwi or bsi.
After running the script 1.1, it is expected a figure like the following:
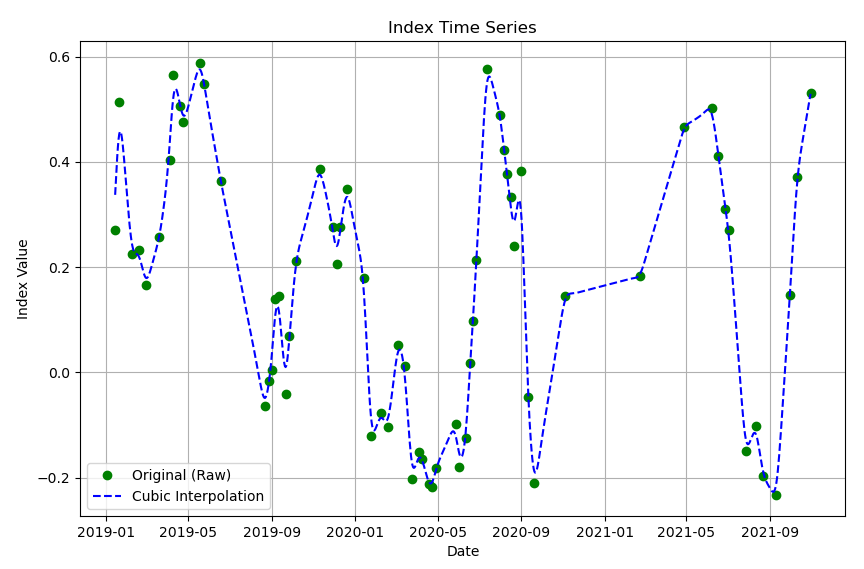
Step 3:
The second script in this lesson, 1.2_imag_reader_generate_smoothed_maps.py, is the smoothing automation for all the pixels. So, for each valid pixel according to the SCL, a smoothed daily timeseries is generated.
The paths for the timely smoothed data are in ./data/interpolated_images/ inside the subfolders bsi and ndwi.
Step 4:
In the sequence, the spatial smoothing is done over the timely generated images with both scripts: 1.3.a_gauss_smoother.py and 1.3.b_gauss_smoother.py. The “a” version of the script generates the NDWI daily images, while the “b” version, the BSI.
At these scripts the smoothing variable (spatial this time) is the sigma_x. Also, if the desired to implement the gaussian smooth to the timeseries, it is possible with sigma_t.
The smoother images generated by both scripts are in the respectively in subfolder bsi and ndwi, which are inside folder: ./data/interpolated_images/_gauss_filt/.
Step 5:
At this step, the ground truth data is transformed into a csv file. The csv file contains the coordinate, four soil layers for each profile (the fourth is the repetition of the third), the local elevation, the texture and the SOC. The bulk density was initially calculated based on the texture and the SOC of the soil sample; the theory is described in the scientific article written by Hollis et al. (2012). A Hollis implementation is available at the script named hollis_bd.py, inside the folder: ./py_codes/libs.
The output csv is written in: ./data/true_data/selection_GIS_32633_ws.csv.
In the following table, a description of the content of selection_GIS_32633_ws.csv file.
| Field | Name |
|---|---|
| Field_name | Name of the field |
| Field_Profile_number | Profile number |
| Horizon_number | horizon number |
| Easting | Longitude (EPSG 32633) |
| Northing | Latitude (EPSG 32633) |
| measured_yield | Yield (tons per ha) |
| Yield_kg_per_ha | Yield (kg per ha) |
| height | local elevation |
| SAND | Sand content |
| SILT | Silt contant |
| CLAY | Clay content |
| Upper_depth | Upper depth of the soil layer |
| Lower_depth | Lower depth of the soil layer |
| Thickness | Thickness of the soil layer |
| KA5 | German soil classification |
| SOC | Soil organic carbon content |
| BD | Bulk density in g/cm3 |
This exercise demonstrates the DSM workflow. The whole map is already provided, and we will simulate that we are not aware of the data. At this step, the script just fetched the data from the file ./data/true_data/soil_Boo_lean_all.csv, which already contains all the data!
Step 6:
A brief analysis of the soil data available in file selection_GIS_32633_ws.csv, shows that the soil texture is kept almost constant, while the SOC shows a higher variability through different layers of the same soil profile.
To show this characteristic, the following table, containing the soil profile data from profile number 848, and the statistics of the Sand, Clay and SOC, is presented.
Table 1 – Statistics of the soil data from soil profile number 848: the mean, standard deviation, and the coefficient of variation (CV).
| Layer no. | Sand [%] | Clay [%] | SOC [%] |
|---|---|---|---|
| 1 | 85.857 | 3.500 | 0.728 |
| 2 | 84.763 | 3.568 | 0.267 |
| 3 | 86.562 | 3.579 | 0.382 |
| 4 | 86.562 | 3.579 | 0.382 |
| mean | 85.936 | 3.556 | 0.439 |
| deviation | 0.736 | 0.033 | 0.173 |
| CV | 0.009 | 0.009 | 0.393 |
The coefficient of variation (CV) is defined as the standard deviation divided by its mean value. The CV can be understood as a normalization of the standard deviation, allowing the direct comparison of different quantities, such as sand, clay and SOC. The CV analysis shows that the SOC is the most important variable related to the different layers of the same profile.
Although Table 1 shows only one profile, the attendant is invited to calculate the CV of the soil for the other selected locations.
Step 7:
The Gaussian filter was not explained in detail as the Whittaker smoothing technique. We recommend the attendant to briefly search and understand that it’s the foundation.
Tasks
- Navigate through the downloaded files and feel comfortable with its folder structure.
- Study the Whittaker smoothing method. It is a technique generally used to smooth a data series. Why are we using it to interpolate?